BioByte 022: human immunome project, male-derived egg cells, dual use of AI in drug discovery, are neurodegenerative diseases really just autoimmune diseases, and more
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. If you’d like to connect or collaborate, please shoot us a note here. Happy decoding!
A huge thank you to everyone who filled out the survey and left us with many comments and thoughts on how to make Decoding Bio better. We’ve read every word and are working on bringing you more of what you want!
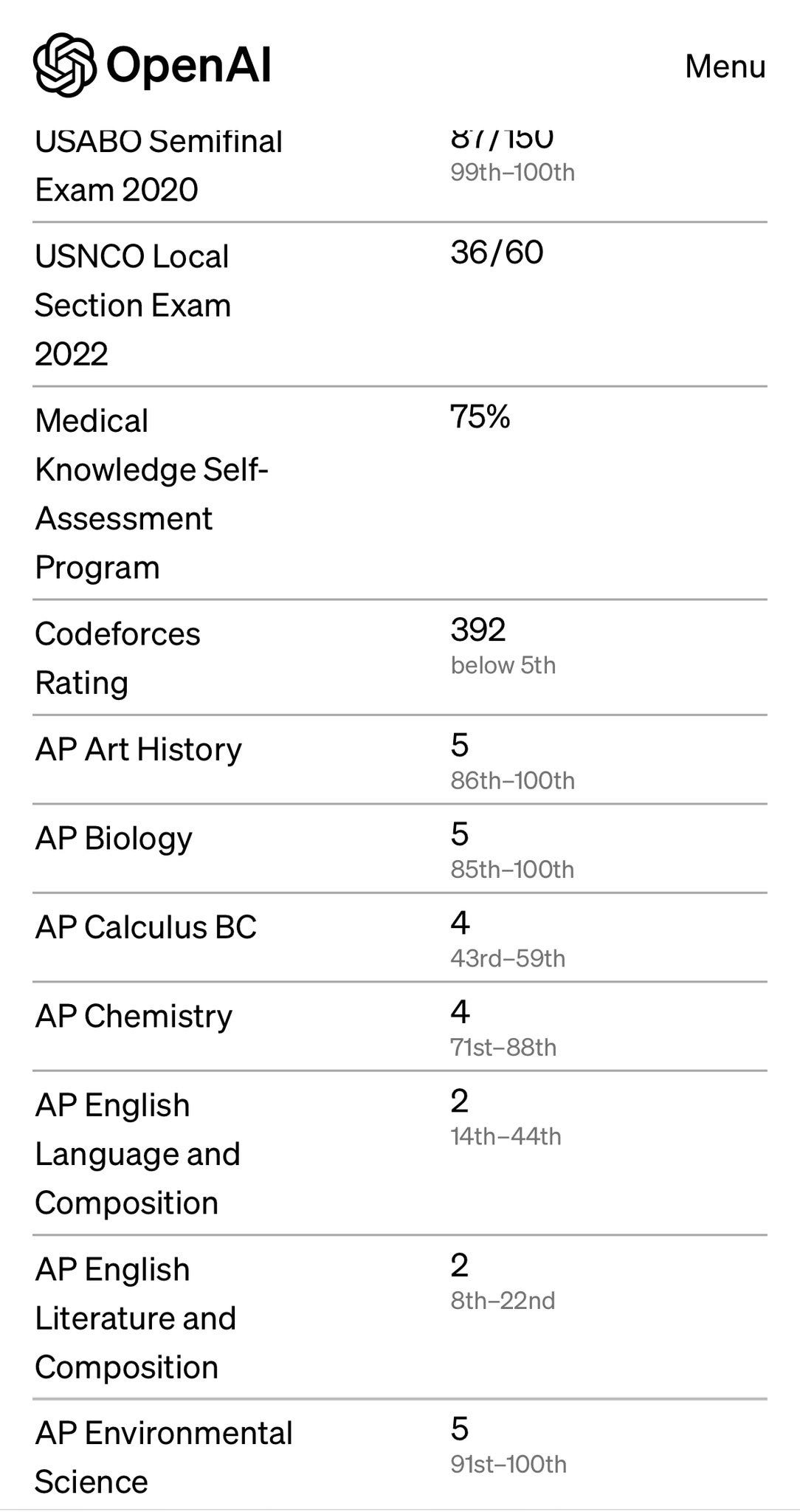
In other news, GPT-4 scored a 5 on the AP Biology Exam and a 4 on the AP Chemistry exam. While this certainly speaks to the power of GPT-4, it also touches on what these exams are testing for. For anyone who has taken the AP Biology exam, you know how much rote memorization it requires. I recently came across this post by James Somers on why he should have loved biology. It’s a beautiful read that highlights the injustice of how biology is taught in schools. Rather than focusing on the mysteries, hypotheses, and principles of life, we teach conclusions of studies as if they are fact. In James’ words:
In the textbooks, astonishing facts were presented without astonishment. Someone probably told me that every cell in my body has the same DNA. But no one shook me by the shoulders, saying how crazy that was.
We’ll drop it in Field Trip but highly recommend James’ inspiring post on how biology should be presented. On to this week’s reads!
What we read
Blogs
The Human Immunome Project is the Next Big Leap [Jane Metcalfe, Proto.Life, 2022]
In the next five years, The Human Immunome Project aims to build a quantitative and predictive model that will improve our understanding of the interactions of immune cells and their components over our lifetime, their interaction with the environment (called the exposome), and how this varies from person to person. The model will be able to anticipate how an immune system will respond regardless of whether the mechanism behind the response is understood or not.
Jane, who is chair of the board, describes a few reasons why this is a complex issue to solve:
Academic silos: researchers in distinct disciplines don’t interact with each other
Data silos: even when experts speak to each other, the data they have gathered will be inaccessible to each other. There is little to no incentive to share data, and many bureaucratic barriers exist that prevent doing so.
Data insight gap: researchers generate data using the latest technologies, but analyze it using yesterday’s methods. Often this data is not a machine-readable format, and when it is there are systematic differences which make the data vary lab to lab and machine to machine.
Magnitude of data capture: huge dimensionality of longitudinal data gathering across the ‘omic stack, in different geographies.
While the project will have to overcome several barriers, the outcome is worthwhile. In ten years, the project aims to bring personalized models of each of our immunomes, based on current health state, clinical past and future projections. This could improve prognosis, early detection and faster interventions.
If you have access to immunological data and want to support the project, you can find out more here.
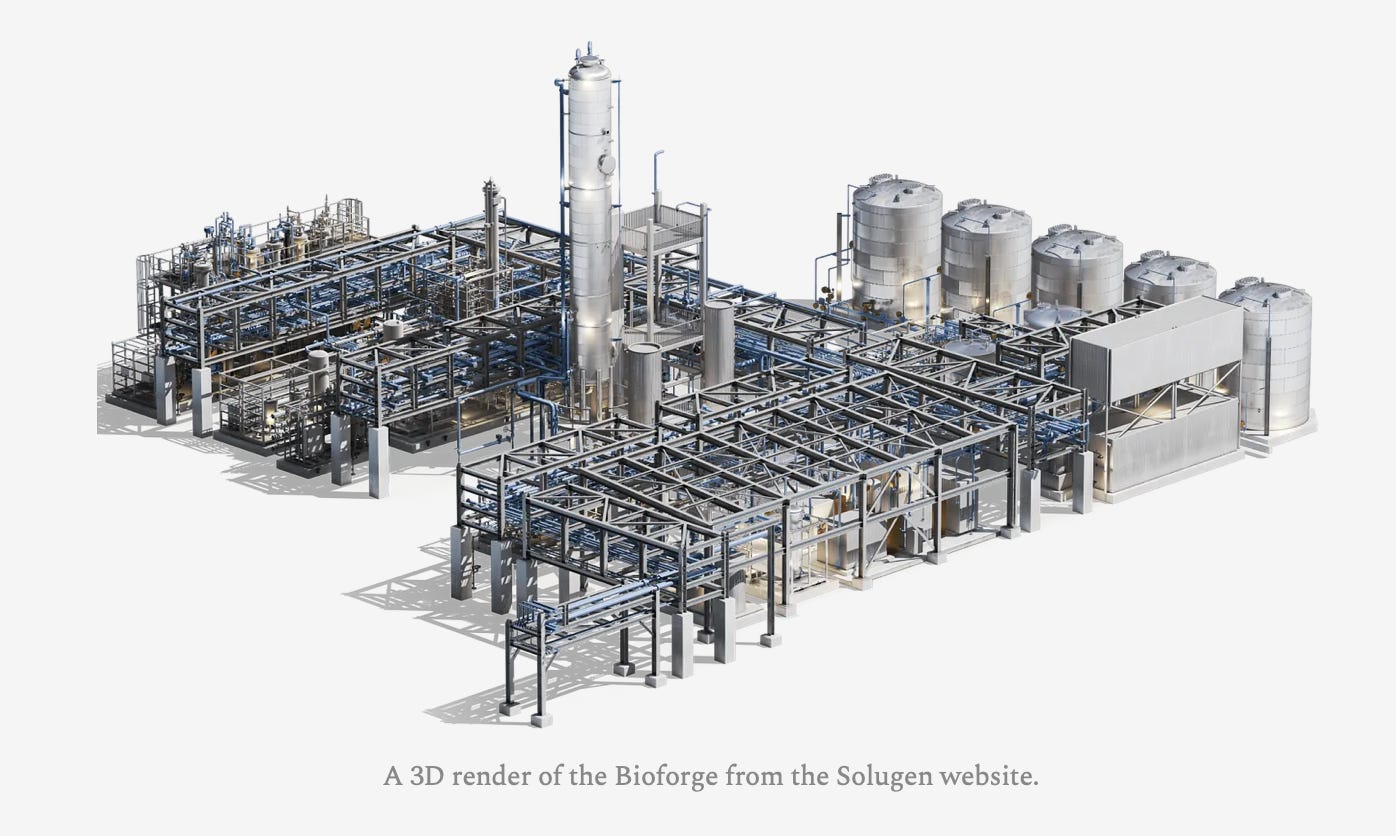
Solugen [Elliot Hershberg, The Century of Biology, 2023]
Friend of DB, Elliot Hershberg, is back this week with a fascinating piece on Solugen. Founded in 2016 by Gaurab Chakrabarti and Sean Hunt, Solugen is a biomanufacturing company that is decarbonizing the chemicals industry. From the techno-economic analysis that proved the feasibility of Solugen’s thesis to delivering Solugen’s first product to customers in Sean’s Subaru, Elliot recounts the early days of the company. Today, Solugen’s Bioforges, or chemical factories that leverage biology, run on sugars, water, catalysts, and air to achieve better synthetic yield than petrochemical or fermentation alternatives, all without using CO2. In the future, Solugen’s goal is to design a library of enzymes that can make 90% of chemicals by 2030 across a distributed network of bioforges. Check out the full piece, it is worth the read. (*Disclaimer: KdT and Cantos are investors in Solugen)
Academic papers
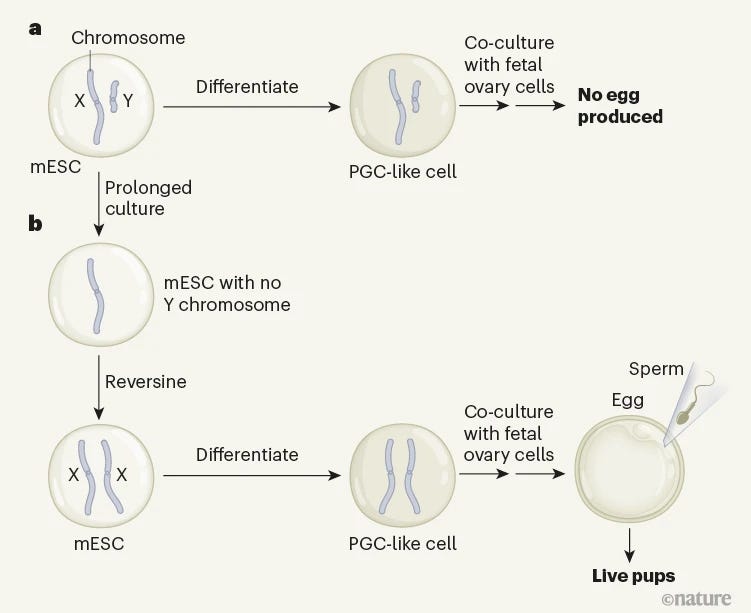
Generation of functional oocytes from male mice in vitro [Murakami et al., Nature, 2023]
Why it matters: We are getting increasingly closer to in vitro gametogenesis, or enabling reproduction through stem cell reprogramming. There have been several studies showing successful transformation of embryonic stem cells to functional oocytes (egg cells) in mice and even studies showing healthy mouse pups being born from such techniques. This paper shows that oocytes can also be created from male mice through chromosomal rearrangement, which opens the door to opportunities for same-sex reproduction and reducing sex-chromosome based disorders. Last week, the Saitou lab published a new paper showing the creation of functional oocytes from male mice. Oocytes, or egg cells, have to come from female (XX) cells, so the scope of the work is quite impressive. The researchers started with embryonic stem cells from male mice and let them divide in culture long enough for spontaneous chromosomal rearrangement to occur. Since genomic material is highly unstable for stem cells in culture, they were able to isolate XY cells that lost the Y chromosome. In fact, 1-3% of XY cells will spontaneously lose the Y chromosome so no synthetic genetic engineering was necessary. To duplicate the X chromosome and get XX, they added the enzyme reversine which increases the probability of chromosomal rearrangement during mitosis. They isolated cells with two XX chromosomes using fluorescent expression and ended up with XX (female) cells that were genetically identical to the initial XY (male) cells. To create oocytes, they used a previously reported procedure involving converting stem cells to primordial germ cells and then maturing into oocytes. Despite the low efficiencies, this protocol shows a clever way to create viable eggs from male starting cells, which has major repercussions for fertility, sex-chromosome based disorders, and same-sex reproduction. These methods also present an opportunity to select against chromosomal-diseases including various trisomies.
Dual use of artificial intelligence powered drug discovery [Urbina et al., Nature, 2022]
Why it matters: Biosecurity is top of mind as biology becomes increasingly programmable with advances in AI and synthesis. This paper raises important questions on the potential dual-use of many technologies developed to design better drugs. Though first published last year, recent advances have us revisiting this paper on the dual use of AI in drug discovery. AI holds much promise in screening compounds and figuring out ways to lower toxicity, increase binding efficiency, and more. However, with the same tools, the inverse is also true—there is potential to design molecules with increased toxicity among other features. This paper explores what would happen if the same tools used to design better drugs are operated by a bad actor for misuse. They used a de novo molecule generator they developed called MegaSyn, normally used to predict bioactivity and find therapeutic inhibitors, to reward toxicity and targeted activity instead of it’s standard penalize toxicity and reward targeted activity. They used public datasets to train the AI and chose to drive the generative model towards nerve agent VX, “one of the most toxic chemical warfare agents developed during the twentieth century”. In six hours, using locally-hosted servers, the researchers found their model generated 40,000 potential toxins, many with similar predicted potency to VX and some with even higher predicted toxicity.
The irony here is not lost on us. Models that were initially built and trained to avoid toxicity in the generation of new drugs were easily modified to generate chemical toxins on par with and more potent than the most deadly of known chemical weapons. Though none of these molecules were synthesized and tested, it’s clear that we need to have an urgent conversation on biosafety, biosecurity, and possible safety controls for AIs that may be turned around for dual use purposes by bad actors or even nonhuman machines.
Microglia-mediated T cell infiltration drives neurodegeneration in tauopathy [Chen et al., Nature, 2023]
Why it matters: Are neurodegenerative diseases like Alzheimer’s actually autoimmune diseases? A new paper in Nature shows that T cells are recruited (possibly by microglia) to the site of tau aggregation and neurodegeneration in mouse models of AD, suggesting that the adaptive immune system may be a promising new therapeutic target. In our 2023 predictions piece, we previewed that it would be an important year for microglia research. This month in Nature, David Holztman’s group at Wash U led by Xiaoying Chen provides new evidence for the role of both the innate (including neuro-resident immune cells like microglia) and adaptive (B and T cells) immune systems in tau-mediated neurodegenerative disease like Alzheimer’s. The neuro-immunology field has come a long way, and quickly; a lymphatic system in the central nervous system was only recently discovered, which overturned many basic assumptions about the role of the immune system in neurological disease.
In this paper, Chen et al. showed that in an ApoE4/tau transgenic mouse model, (primarily) CD8+ cytotoxic T cells are summoned to the brain by microglia and aggregate in areas of tau pathology, and removal of these T cells prevented brain atrophy. In more detail - first, to characterize the immune response in neurodegenerative disease, the authors compared single-cell RNA-seq data of immune cells in amyloid and tau mouse models. Surprisingly, the number of T cells was significantly increased in the tau-aggregation mice relative to the amyloid-deposition mice. Gene expression analysis of T cells showed that effector CD8+ T cells localized primarily to the brain parenchyma rather than the brain border, suggesting that they were responding to an antigen presented by the innate immune system. Single-cell T cell repertoire (TCR) sequencing confirmed T cell clonal enrichment in the parenchyma of tau models of neurodegeneration.
What causes T cell recruitment and activation? Microglia, although the evidence is circumstantial. Microglia expressing MHC class II proteins like IBA1 were significantly increased in regions with neurodegeneration. Gene expression analysis showed that genetic markers for homeostatic microglia were decreased, while genes related to antigen presentation, complement response and cytokines were increased, suggesting an important role for activated microglia in recruiting T cells into the brain parenchyma.
The implications of these findings are important. This is arguably the most compelling evidence yet that neurodegenerative disease like Alzheimer’s is in fact driven by an autoimmune response to pathological protein deposits in the brain. One of the most exciting future directions of this work is characterizing the T cell repertoire (TCR; collection of T cell receptors on the surface of T cells that captures the history of an individual’s antigen exposure) to elucidate specific antigens responsible for driving neuro-pathology, which may result in novel therapeutic strategies. Most drugs for neurodegenerative disease to date have focused on targeting protein deposits or microglia, but very few have targeted T cells.
Decrypting drug actions and protein modifications by dose- and time-resolved proteomics [Zecha et al., Science, 2023]
Why it matters: PTMs are chemical modifications that play a significant role in functional proteomics, as they influence activity, interaction, and localization of proteins. To date, the ability to interrogate PTMs at scale and in high throughput has remained poor. The authors introduce decryptM, a proteomic technique that quantifies drug-PTM modulation, providing the ability to better understand target engagement and mechanisms of action. Post-translational modifications (PTMs) are critical chemical modifications that significantly influence functionality of proteins, with over 200+ unique ones identified. PTMs are significantly affected by therapeutics, and they can provide crucial information about target engagement, phenotypic response, and can identify pathways that are being modulated downstream. Although large-scale proteomics have increasingly come to the forefront of biomedicine, the ability to properly interrogate PTMs has remained poor at best.
Here, Zecha et al. develop an approach called decryptM, in which cells are treated with increasing concentrations of a drug, and are subsequently enriched for PTMs with stable isotopes (tandem mass tags). Following a trypsin digest, subsequent peptides are fractionated via chromatography for proteome profiling, and peptides with PTMs are enriched via immunoprecipitation or chromatography. These dose-dependent treatments are measured together using liquid chromatography tandem mass spectrometry (LC-MS/MS). This provides the ability to obtain dose-response characteristics panned on individual peptides – ultimately informing on target/network engagement.
The authors then applied decryptM to several cancer cell lines x cancer therapeutics (including chemotherapeutics, proteasome inhibitors, epigenetic modulators, kinase inhibitors, antibodies, and more). They identified several interesting things (all of them would be too long for this post), including novel phosphorylation sites that tie chemotherapeutics to DNA damage response pathways, and even found that Rituximab (a very common anti-CD20 antibody, for which a MoA has been poorly understood) kills CD20-positive B-cells by overactivating BCR signaling. You can explore decryptM here!
What we listened to
Notable deals
Switch Therapeutics comes out of stealth with $52M for RNAi therapeutics.
Shennon Bio raises $13M for immune cell profiling
Pfizer acquires Seagen in $43B deal to expand cancer offerings
Artera AI launches with $90M for biomarker-based cancer diagnostics
In case you missed it
What we liked on Twitter








Events
AI + Bio Hackathon by Latch [San Francisco, March 25]
Field Trip
Why I Should Have Loved Biology, James Somers
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone












You write...
"Though none of these molecules were synthesized and tested, it’s clear that we need to have an urgent conversation on biosafety, biosecurity, and possible safety controls for AIs that may be turned around for dual use purposes by bad actors or even nonhuman machines. "
This is a very well intended sentiment which I'm sure is sincere.
But urgent conversations on bio and AI safety are not going to make us safe, because the people we should be worried about will ignore such conversations, rules, laws, governance schemes etc. As you know, much of the planet is ruled by ruthless dictatorships with vast resources at their disposal. And, bio and AI developments will empower smaller evil doers like never before.
Think bigger. Much bigger.
Almost all the violence in the world at every level of society is committed by a small fraction of the human population, violent men. That should be the target. Identify the source of the problem in a clear minded direct manner, and remove it.
If the world of biology can present us with a credible plan for liberating humanity from the lethal infection of violent men, that is a proposal I would consider with extreme interest. Until then, stop doing biology, and start learning how to think bigger. Here's a place to begin perhaps...
https://www.tannytalk.com/p/world-peace-table-of-contents