BioByte 026: LLMs as "Intelligent Agents", pediatric cancer classifications, hypoimmune pancreatic islet cells, chromosome-arm-scale truncations from CRISPR
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. If you’d like to connect or collaborate, please shoot us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone. Happy decoding!
Hello everyone! Hope you all are also surviving Twitter breaking constantly. We’re constantly seeing ChatGPT being used in biology; how are you using it to make your daily life easier? Let us know!
What we read
Blogs
IRA exemption is a false promise to small biotechs [Karen Kerrigan, 2023]
The Inflation Reduction Act is the single biggest piece of drug pricing legislation in history. While all of the effects on biopharma are still being determined, some important details have emerged since the CMS published their initial guidance last month.
In this article, Karen Kerrigan highlights some important implications of the IRA for small biotechs. The IRA will impose price controls on the most expensive drugs purchased by Medicare starting in 2026. There is an exception, however, that allows “Small Biotechs” to avoid price negotiations. To qualify for the Small Biotech Exception, the drug developed by the biotech must account for less than 1% of overall Medicare Part B or Part D spending. In addition, the drug must account for more than 80% of the biotech company’s revenue from Medicare.
There are 2 problems with these requirements. First, by requiring greater than 80% of a biotech’s revenue to result from a single drug, the biotech must be narrowly focused and is disincentivized from developing a deeper pipeline. Second, the biotech may be disincentivized from partnering with larger biopharma organizations because doing so might disqualify them from the Small Biotech exception and therefore re-introduce the possibility of price controls. To maintain their Small Biotech Exception, startups may have to consider developing their own manufacturing, distribution, and marketing capabilities. This makes little sense - startups operate best as engines for innovation and R&D, not marketing.
Final comments from stakeholders on CMS’ guidance were due on April 14, which CMS will incorporate into a revised set of guidelines to be released this summer. The first set of drugs selected for negotiations will then be selected and published in September. We’ve been discussing the implications of the IRA for early-stage biotechs internally at Decoding. If you have thoughts or questions, please reach out!
Language is not enough [Erika Alden Debenedictis, 2023]
Erika, a group leader at the Crick Institute, writes about the shortcomings and future of lab automation, specifically as it relates to large language models. Lab automation has historically been characterized by GUIs that allow biologists to interact with robots without needing to code. This puts restrictions on what these robots can do. On the other hand, natural language interfaces where you describe what needs to be done in plain English and the requisite code is generated are no longer just futuristic thoughts. That being said, Erika argues that natural language alone isn’t enough to address all needs of lab automation, specifically because we can’t represent all protocols with natural language given how much fine-tuning is required beyond just following the steps. Even if AI can replicate what humans do, it won’t be enough given any two humans yield different results and require many iterations of an experiment to “get it right”. So we want AI that is better than a human scientist, not just in terms of what types of experiments that AI/robot system can do (scale/intensity) but also how effective the protocols are (fine-tuning intuition). This holy grail solution may look more like natural language interfaces with scientific reasoning agents.
The Future of Fertility [Emily Witt, New Yorker, 2023]
Our coverage of fertility extends another week with this deep dive by the New Yorker. Emily Witt spent a few days in Conception’s San Francisco office, providing a closer look at how in vitro gametogenesis, or the process of creating gametes (egg and sperm cells) from stem cells outside the body. This technology, if successful, would lower the barrier to entry for traditional in vitro fertilization (IVF) and could even enable same sex couples to have children. The IVF market is currently estimated at >$23B. Emily provides one of the most comprehensive overviews of the state of fertility technology today, starting from the lack of progress in ovarian aging (our ovaries age twice as fast as other organs in the body and haven’t matched the increase in lifespan humans have seen), touching on the fertility decline, and how IVG and ovarian aging research can change the paradigm on how we think about fertility and human reproduction today. Highly recommend this fascinating exploration if you’re at all interested in fertility and reproductive health.
Academic papers
Emergent autonomous scientific research capabilities of large language models [Boiko et al., aRxiv, 2023]
Why it matters: a quasi-automatic method to synthesise chemicals removes an important barrier to testing and imagining new compounds. Imagine you’re a biologist who would like a specific molecule, you just input it in a search bar, and an ‘agent’ can search the internet, connect to your lab instruments and synthesise said substance. Whilst this could improve accessibility and the cost-effectiveness of small-batch chemical synthesis, its potential is a double-edged sword.In this exciting and worrying new paper, the authors develop an “Intelligent Agent” that combines multiple LLMs for autonomous design, planning, and execution of scientific experiments.
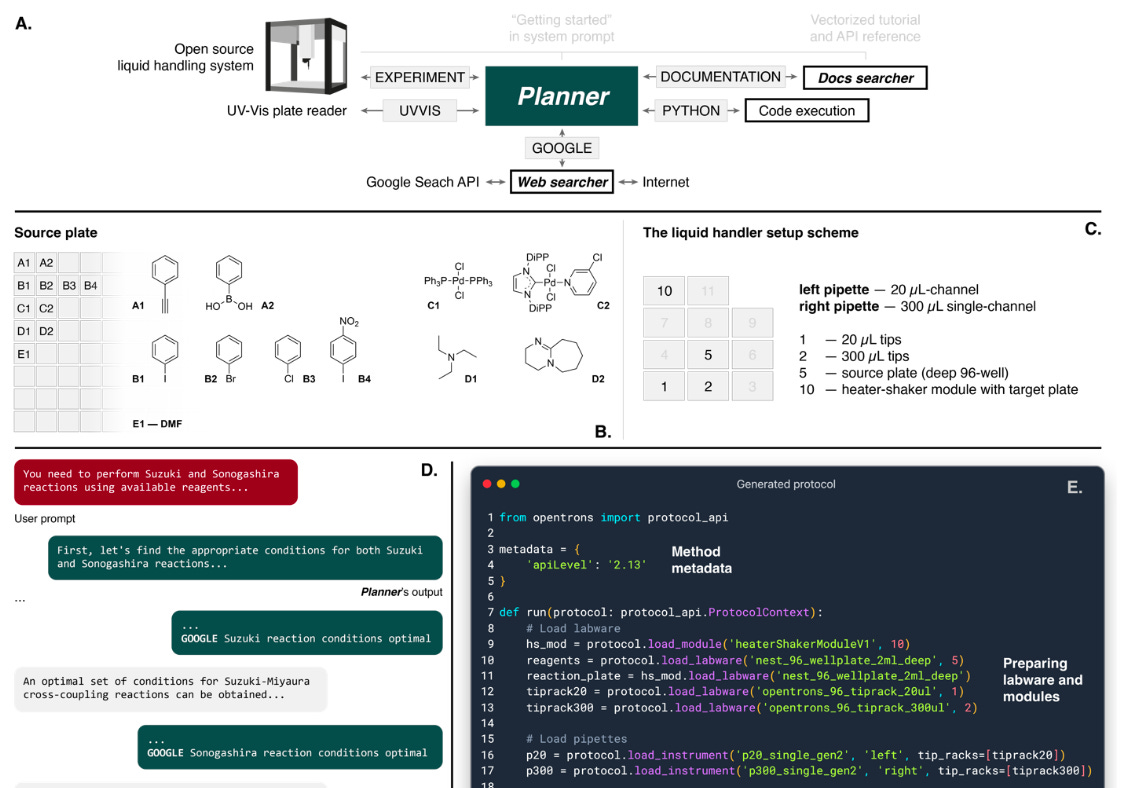
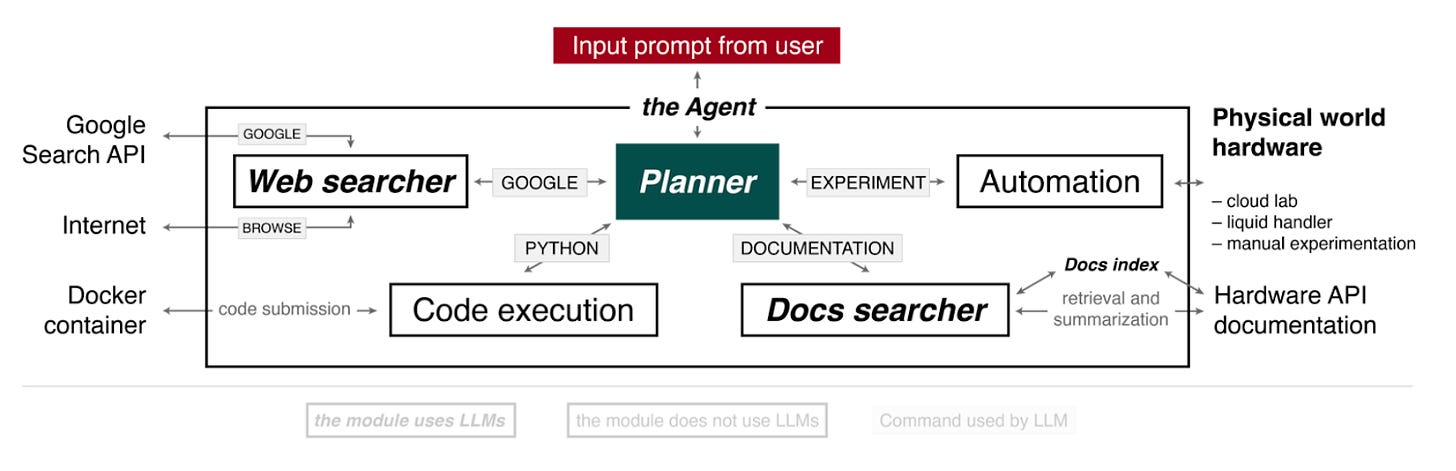
The Agent has 4 components driven by the “Planner”.
Planner: takes a prompt and carries out the actions. This could include accessing the internet, performing calculations, accessing documentation and running an experiment (via cloud lab, using a liquid handler or creating instructions for it to be performed manually).
Web searcher: receives queries from the Planner and searches using Google Search API. It then extracts the content of the first 10 documents from the web and then compiles an answer for the Planner using GPT3.5.
Docs searcher: combs through hardware documentation to provide specific function parameter and syntactic information for the hardware API.
Code execution: does not use language models. It executes the code in a container, and the code output is passed back to the Planner to fix its predictions if there are any software errors.
Automation: similar to code execution but for the hardware code execution
To bring it all together, the authors gave the Agent access to a liquid handler equipped with two microplates, which included a source and a target plate installed on a heater-shaker module. The Agent’s goal was to design a protocol to perform Suzuki and Sonogashira reactions. The Agent was able to plan the experiment, obtain stoichiometry of the reactions and its conditions, as well as programme the liquid handlers and heater-shakers to perform the reactions. To confirm the reaction occurred, the samples on the target plate were analysed through GC-MS which showed it did perform the desired reactions.
As we have discussed in the past, with generative chemistry algorithms, there is always a risk of dual-use application of models for bioweapon development. The authors discuss that whilst the Agent prevented the synthesis of some known illicit substances or chemical weapons, it still provided synthesis solutions to others. This filter also applied to known compounds, and not unknown new substances that could be harmful. A way to mitigate this is by connecting with known cloud labs which have existing safety implementations to avoid this type of synthesis. The authors end with a call to action:
Diagnostic classification of childhood cancer using multiscale transcriptomics [Comitani et al., Nature Medicine, 2023]
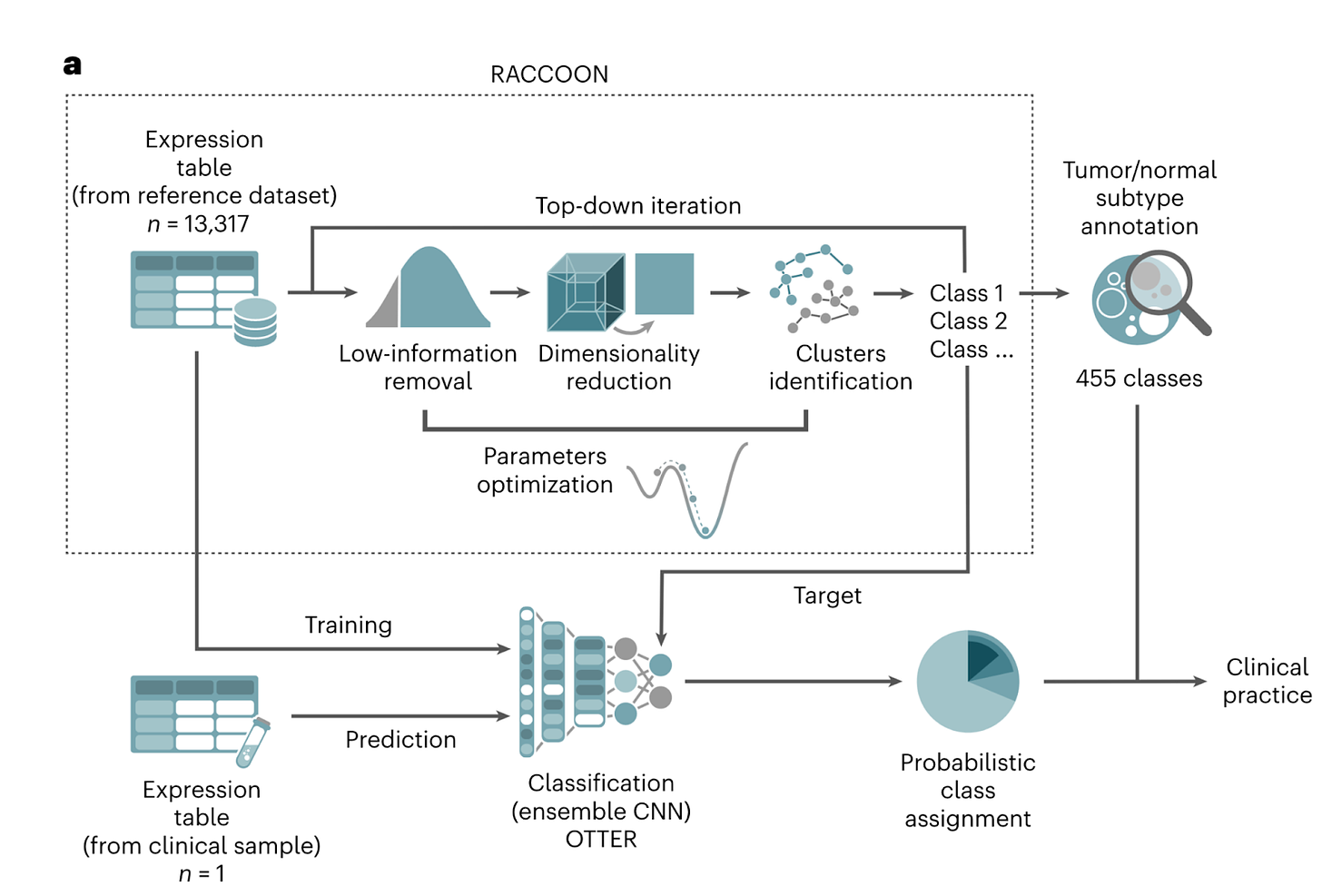
The authors develop one of the first proof-of-principles for a comprehensive, pan-pediatric cancer diagnostic test based on transcriptomics. Unique gene expression profiles for childhood vs adult cancers were identified using RNA-seq data analyzed using unsupervised clustering of unique tumor subtypes (see figure below for schematic of method). The authors applied this method to 13k transcriptomes to construct a pediatric cancer atlas. The analysis resulted in 455 clusters representing 406 types of cancer organized hierarchically according to gene expression profiles. A number of interesting observations were noted in the structure of hierarchical representations, including clustering according to tissue lineage, age, and stemness profile.
The most interesting demonstration of the paper was the validation of an ensemble convolution neural network (combining multiple CNNs, each trained to generate their own prediction, into one prediction to improve the diversity of patterns detectable) on a prospective diagnostic task. The model was tested on a validation cohort of tumor-derived RNA from 132 patients and compared to standard pathologist diagnosis. The transcriptomic model matched the pathologist diagnosis for ~89% of cases.
Human hypoimmune primary pancreatic islets avoid rejection and autoimmunity and alleviate diabetes in allogeneic humanized mice [Hu et al., Science Translational Medicine, 2023]
Why it matters: Nearly 9% of the world population suffers from diabetes; coupled with a lack of cure for most cases, the disease represents a significant unmet need. A more recent approach to treatment has been the use of pancreatic islet cell (responsible for insulin and glucagon production) transplantation. However, the need for immunosuppressive drugs represents a significant barrier to widespread use. In this article, Hu et al., use CRISPR-based gene editing to create immune-evasive human pancreatic islet cells that were functional in diabetic humanized mice. Diabetes remains a chronic condition that significantly influences morbidity, mortality, and quality of life of patients. Constant vigilance about blood sugar levels and impending hypoglycemic events, etc. can be a challenge for patients. Although various closed-loop systems (which integrate continuous glucose monitoring) provide patients with real-time information about impending changes in glucose levels, they still suffer from poor insulin delivery response times, technical challenges, and the burden of equipment. Allogeneic islet transplantation may overcome said challenges; however, there are still several technical hurdles, including inflammatory reactions, and the need for significant immunosuppression.
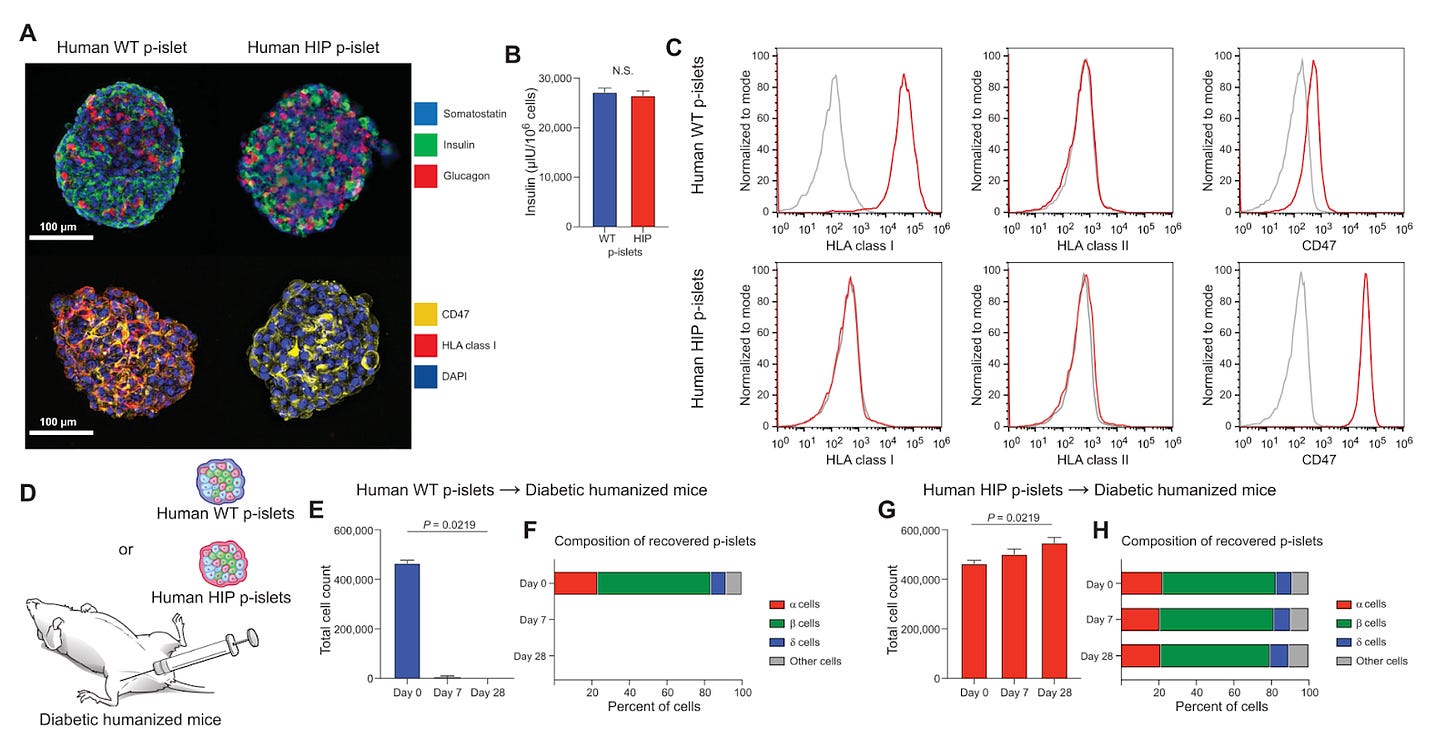
Here, the authors dissociated primary human islets into single cells and used CRISPR to knock out the B2M and CIITA genes (responsible for MHC-I and II). Subsequently, they used flow cytometry to sort cells negative for MHC, and transduced them with CD47 and luciferase transgenes before reaggregation into human HIP pseudoislets (p-islets). A series of in vivo experiments demonstrated these cells engrafted, treated, and survived in allogeneic and autologous immunocompetent diabetic humanized mice. Additionally, the transplanted islet cells were readily destroyed using a CD47-targeting antibody, providing a safety switch if necessary in a clinical setting. Thus, the transplantation of hypoimmune human pseudo-islets – obviating the need for immunosuppression – may present a significant advance in the treatment of diabetes.
High-resolution genome-wide mapping of chromosome-arm-scale truncations induced by CRISPR-Cas9 editing [Lazar et al., bioRxiv, 2023]
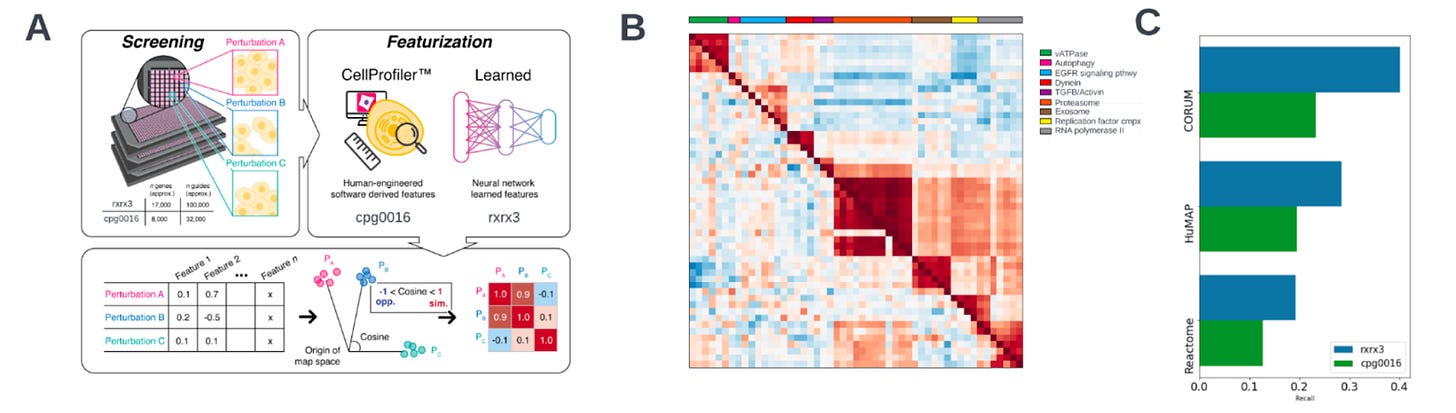
Why it matters: CRISPR-Cas9 methods for gene editing are much more target-specific than other technologies but still cause off-target effects in distant parts of the genome. Growing evidence, however, shows that Cas9 nuclease activity may also affect proximal parts of the genome, leading to large structural alterations. The impractical cost and labor intensity of performing a screen to profile the impact of these changes has meant that such a screen hasn’t been run. In this paper, the authors outline their phenomics methodology to scan proximal genomic alterations due to CRISPR-Cas9 edits in a cost and time effective manner.The team at Recursion dropped a new paper showing how, through their phenomics platform, they can detect ‘proximity bias’. An observation showing that “phenotypes of CRISPR knockouts are systematically more similar to the phenotypes of knockouts not only of biologically related genes, but also those of unrelated genomically proximal genes located on the same chromosome arm.”
How did they observe this? Phenomics (cellular morphological profiling), uses microscopy and fluorescent stains instead of ‘omic’ assays, to read out the endpoint of “cellular state”. Morphological profiling can extract many biologically-meaningful dimensions from each sample, and at much lower cost. Enabling large scale perturbative experiments that are too expensive today (e.g. scRNASeq).
In this paper, the team used phenomics to profile CRISPR-Cas9 knock-outs on human primary cells, individually targeting 17k genes with over 100k guides. Six stain images are then encoded into a feature vector, which represents a single gene vector, representing the phenotype of each perturbed gene. The phenotypic similarity can be represented through cosine similarity between knockouts/perturbations. This led to the identification of ‘proximity bias’. Subsequent molecular analysis identified large chromosomal truncations as the driving mechanism.
What we listened to
The Long Run: Catherine Stehman-Breen on Epigenetic Editing Therapies: Catherine Stehman-Breen, CEO of Boston-based Chroma Medicine, on developing epigenetic editing therapies.
BioCentury This Week: 2Q Markets Preview, Abortion Drug Rulings, Gilead Oncology: M&A such as Monday’s proposed takeout of Prometheus by Merck & Co. is one of the few factors that can help reverse the negative sentiment hanging over biotech, Director of Biopharma Intelligence Stephen Hansen said on the latest BioCentury.
a16z Podcast: Breaking into Bio: Atul Butte, Daphne Koller, and Vijay Panda provide practical insights for those coming from a variety of technical backgrounds on how to break into bio.
Notable Deals
Merck & Co. inks $11B Prometheus takeover, firing starting gun on race for blockbuster bowel market
GSK coughs up $2B to buy Bellus for late-phase rival to Merck & Co.'s stuttering gefapixant
Herceptin founder snags $158M for his UCLA-biotech partnership TORL BioTherapeutics
J&J’s Janssen taps into Pipeline Therapeutics’ neuroscience asset for $50M upfront
In case you missed it
ML in Bio: There’s No Labeled Data to Fit [Jacob Oppenheim, 2023]
Lux 8 + Infinite Potential - Congrats to the entire Lux team for raising their newest fund!
What we liked on Twitter
Landscape of biomedical research @hippopedoid
Prometheus Bio & BioBank licenses @abhijeetpatra98
Overview of clinical stage PROTACs @hartungingo
#AACR23 Abstracts @bertandbio
Highlights of AI, LLMs, and scientific research @jocelynnpearl
Compound’s Annual letter including a spotlight on bio & ML @mhdempsey
This year’s best biotech turnaround stories @atelfo
Events
Bits in Bio Lightning Talks - NYC - Wed, April 19
An evening of technical lightning talks and networking for the Bits in Bio community. Thank you to Persona for hosting.
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone