BioByte 035: LLMs and biosecurity, AI-based design of PROTAC linkers, lessons in leadership and management in pharma R&D, transforming the dark chemical space of nature
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. If you’d like to connect or collaborate, please shoot us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone. Happy decoding!
Quick rundown of what’s covered this week:
A specialized office within the FDA for cell and gene therapy regulation
Modeling human embryonic development using stem cells with applications in cancer and antimicrobial therapies
A positive PhI readout for an ALS candidate from Verge Genomics, discovered in part using AI
Lessons in leadership and management in pharma R&D
An ML method for engineering linker domains for PROTACs using reinforcement learning
A self-supervised transformer model for predicting the structure of unknown metabolites
What we read
Blogs
FDA Braces for Looming Boom in Cell and Gene Therapy Submissions [Ana Mulero, Biospace, June 2023]
The field of cell and gene therapies is experiencing unprecedented growth, evidenced by a surge in regulatory submissions. This trend is prompting the establishment of a specialized office at the FDA: the Office of Therapeutic Products (OTP), which was created earlier this year to streamline workflow processes and support the evaluation of cell and gene therapy applications. With the pipeline for these therapies rapidly expanding, industry experts predict a significant increase in regulatory decisions in the coming years.
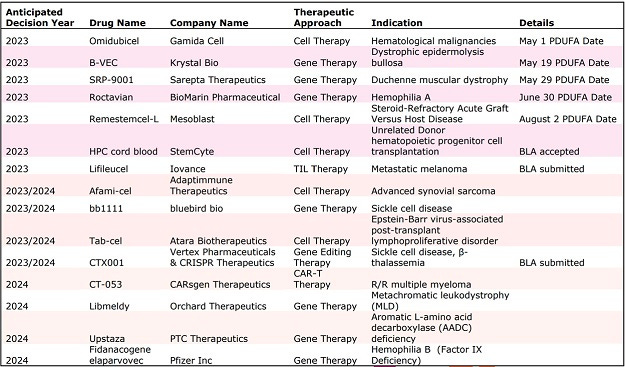
A list of some of the upcoming cell and gene therapies that the FDA might be reviewing/Alliance for Regenerative Medicine:
The FDA's preparedness to handle the influx of submissions has been a topic of discussion within the industry. While the agency is committed to evaluating investigational new drug (IND) applications and ensuring the benefits of proposed studies outweigh potential risks, it recognizes the need to update the regulatory framework to address the unique challenges posed by cell and gene therapies. The establishment of the OTP is a positive step forward, but experts emphasize the importance of modernizing the regulatory paradigm to keep up with the evolving landscape of these therapies.
As the number of cell and gene therapy candidates continues to rise, stakeholders underscore the necessity of standardized processes, improved collaboration, and updated chemistry, manufacturing, and controls (CMC) frameworks. The FDA acknowledges these concerns and is committed to working with industry stakeholders to enhance the efficiency of development and approval processes. While changes will take time, they are crucial to ensure patients can access life-saving treatments in a timely manner. Overall, the establishment of the OTP and the ongoing efforts to modernize regulations reflect the FDA's recognition of the significance of cell and gene therapies in shaping the future of medicine.
Organoids meet single-cell and spatial omics [Yiming Chao, Nature Bioengineering Community, June 2023]
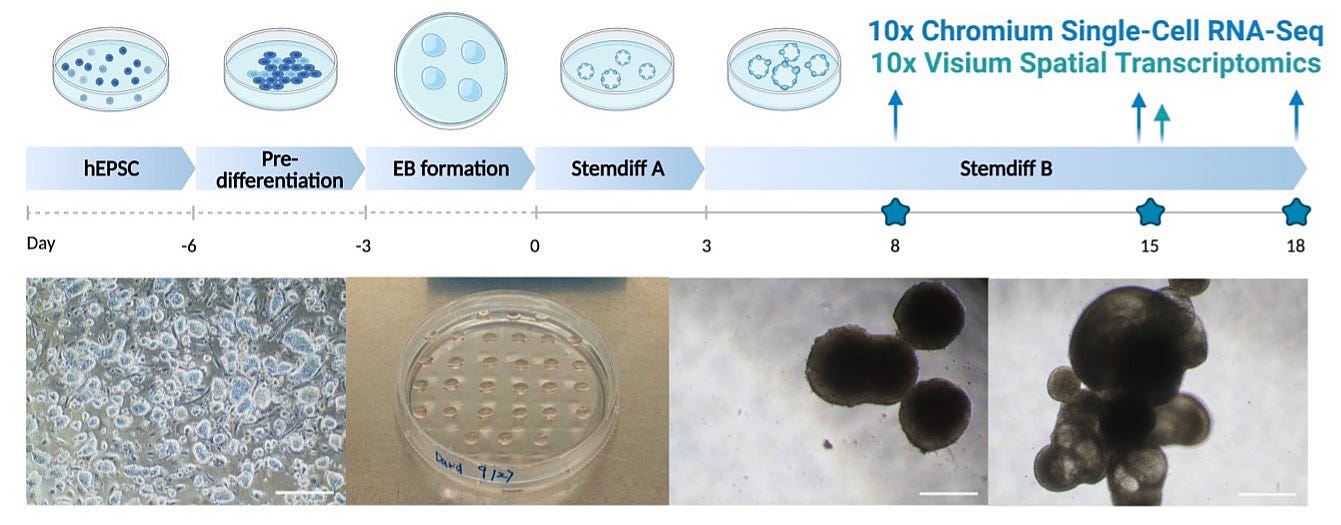
Scientists have made significant progress in modeling human embryonic development in the laboratory using stem cells. A recent paper focused on creating Human Embryonic Organoids (HEMOs) by harnessing the self-organizing capacity of expanded potential stem cells. The HEMOs successfully recapitulated key events in human embryos, including the development of cardiac precursors, neural crest cells, blood cells, placenta, and yolk sac. Through advanced sequencing techniques, the study shed light on cell-cell interactions and gene expression patterns during embryonic tissue specification.
One important finding was the role of trophoblast-like tissues in promoting the maturation and migration of neural crest cells. Additionally, the study identified a yolk sac hematopoietic niche where physical cell-cell interactions occurred between yolk sac endoderm, erythroid cells, and megakaryocytes. The researchers discovered that vitronectin-integrin signaling played a crucial role in promoting megakaryopoiesis in HEMOs, opening up possibilities for generating platelets and immune cells from stem cells for therapeutic purposes.
The ability to model human embryonic development using HEMOs provides a valuable tool for studying developmental biology, engineering genes relevant to human development, and disease modeling. The findings also have implications for the generation and engineering of hematopoietic and immune cells for potential applications in cancer treatment and infectious disease therapies.
Genomic medicines: the coming waves? [Xie et al., Nature Reviews Drug Discovery, June 2023]
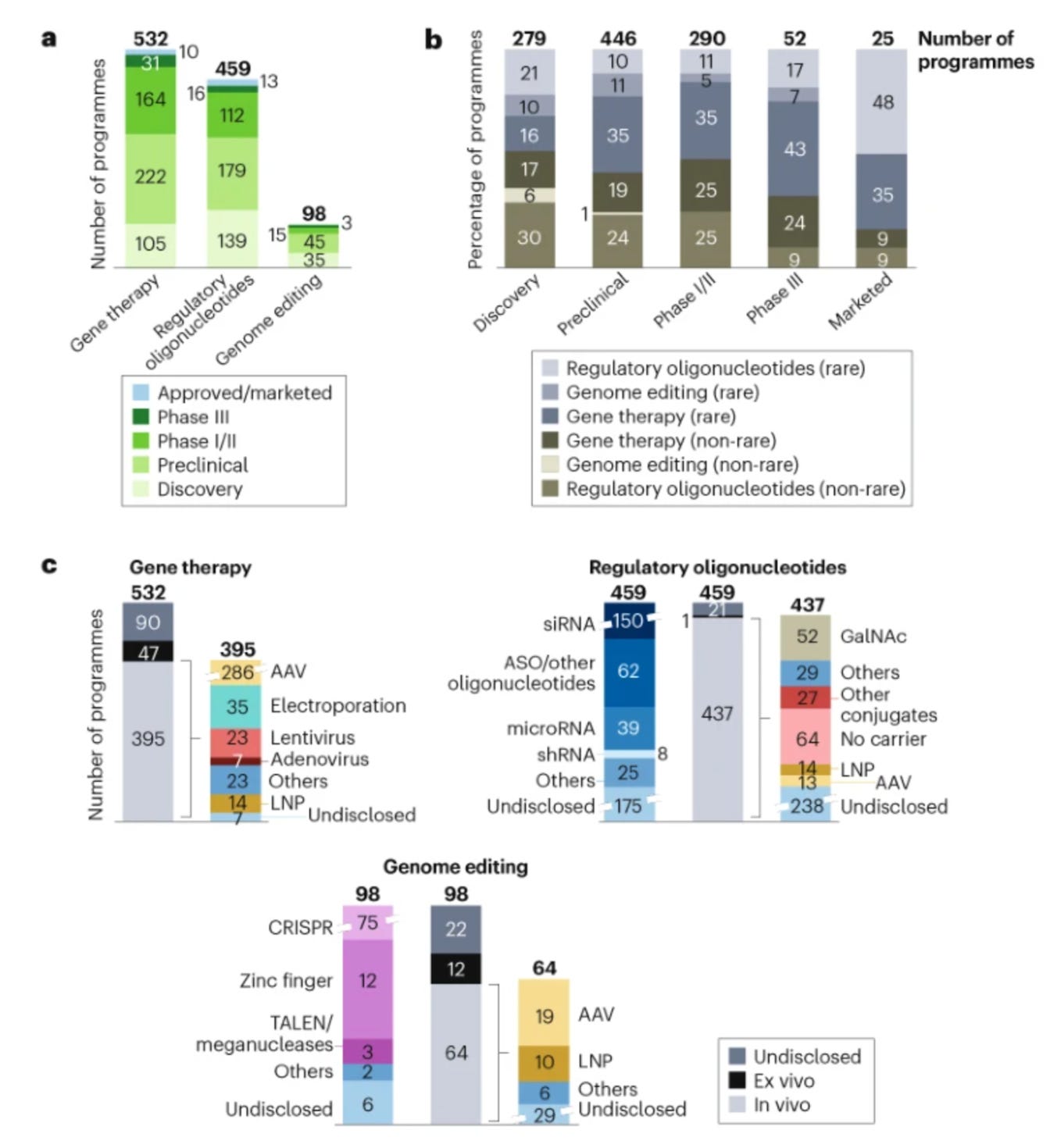
Xie et al analyze the current state of genomic medicine in a few excellent figures. Check out the supplementary info here as well.
Current landscape: there are around 1200 active clinical trials of genomic medicines. Fewer than 20 genomic medicines have been approved so far. Together, they target <0.1% of genetic disorders. 80% of marketed products treat rare diseases.
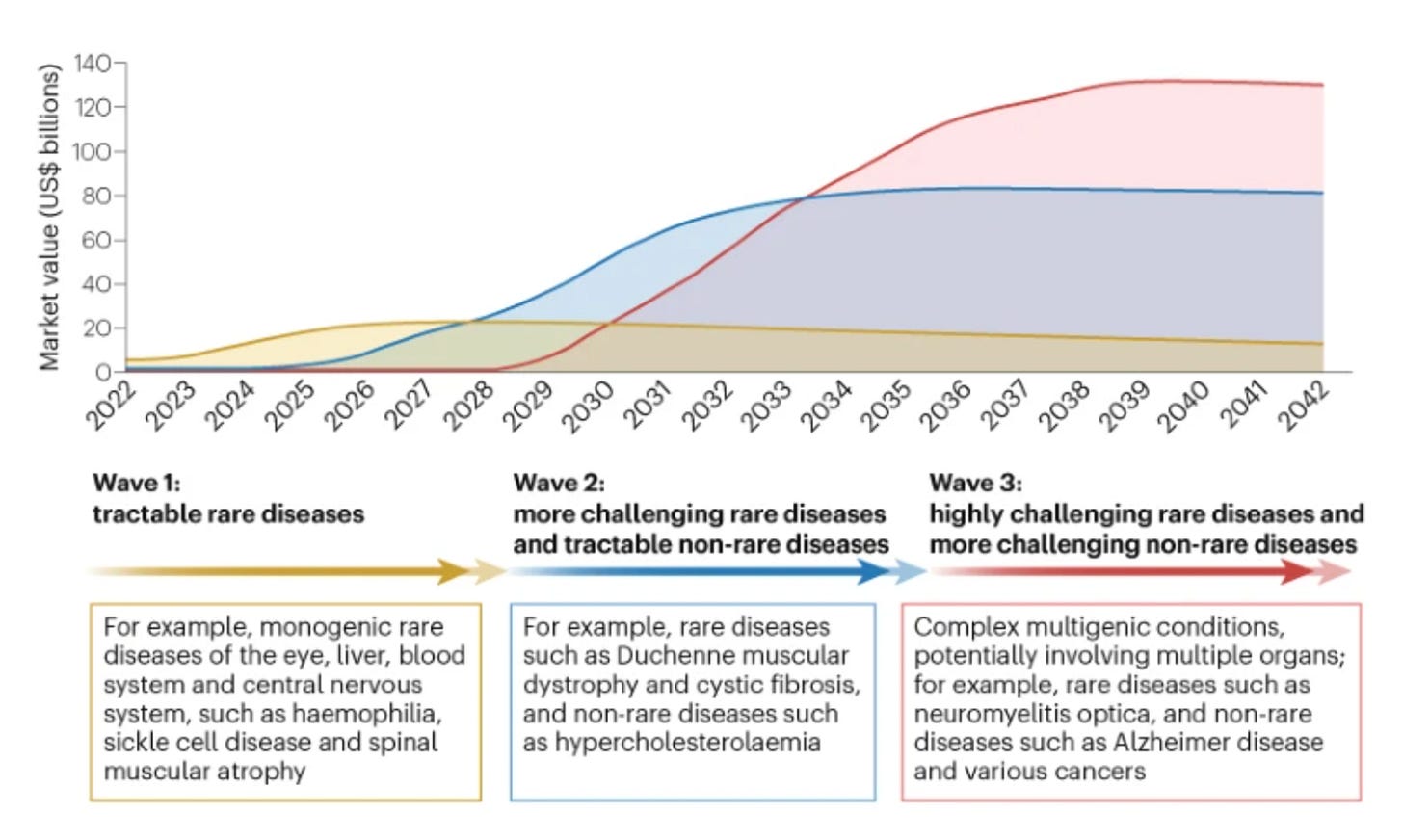
Emerging waves: the authors anticipate three waves of genomic medicines driven by therapeutic tractability of different diseases and advances in targeted delivery:
Verge Genomics shows promising phase 1 results for ALS treatment [Irina Bilous Biopharma Trend, June 2023]
You might remember back in November we discussed Verge Genomics dosing its first patient with its PIKfyve inhibitor therapy for ALS discovered in part by AI. It was a major milestone for AI-enabled drug discovery and we were all cautiously optimistic for what the results might be…and what spell they might cast over a field that desperately seeks clinical validation. Well some of that wait is over. The team concluded their Phase 1 clinical trial of VRG50635 and reported some of the results earlier this week. It’s good news: the drug was well tolerated and demonstrated safety in healthy adults. There were no serious adverse events reported and pharmacokinetics data suggests that once-a-day administration is acceptable. The trial was conducted randomized, double-blind, and placebo-controlled in 80 healthy adults. Verge’s next step is to take back to the clinic at the end of the year for a proof of concept study in patients with ALS. A more telling hurdle, it’s back to the waiting game.
Successful pharmaceutical discovery: Paul Janssen’s concept of drug research [Paul Lewi and Adam Smith, September 2007]
Paul Janssen, as in Janssen the late founder of Janssen Pharmaceuticals (now a subsidiary of Johnson & Johnson), is widely hailed as a renowned and respected drug researcher. The authors of this essay recount some of Janssen’s drug development and management wisdom, straying away from the emphasis on hierarchical management governing decisions back to the individual researcher—a system they claim leads to more breakthrough research. Some highlights of the principles that led to Janssen’s development of 79 novel drugs in 40 years:
People-oriented management as opposed to process-driven allows for maximum breakthrough because individual researchers are allowed the freedoms they need to try things.
Center research around competent people, not rigid processes. This means the organization adapts to the people instead of the people adapting to the organization. Easier said than done, Janssen’s approach questions much of traditional corporate culture. With this style of organization, the company has very little hierarchy between roles and reporting levels. When the organization has to grow, separate teams, or “specialized research units” should be created
Set the research goals based on continuous critical questioning. There should be room for unplanned discoveries made through natural inquisition and creativity.
Leaders should draw ideas out of staff and be an information router. They should be readily available for collaborators. Directors are more like conductors drawing a symphony out of the orchestra.
‘Act as if your own money were at stake’
Keep an open mind—serendipitous findings are often more important than preconceived experiments. Assume as little as possible.
Academic papers
Reinforcement Learning-Driven Linker Design via Fast Attention-based Point Cloud Alignment [Neeser et al., arXiv, June 2023]
Why it matters: In a post-ChatGPT era, we’re all wondering what the most impactful applications of reinforcement learning with human/biological/chemical feedback will by for science. Most recently, scientists have developed a new method called ShapeLinker for designing linkers in Proteolysis-Targeting Chimeras (PROTACs), which are small molecules designed to promote the degradation of disease-relevant proteins. The system leverages reinforcement learning on an autoregressive SMILES generator.Historically, the design of the linker domain in PROTACs has been challenging. The linker domain is a connecting region that physically joins two protein-binding domains. One domain binds to the disease-relevant protein of interest, while the other domain interacts with the E3 ligase. Here is a great overview of PROTAC technology:
Researchers from VantAI propose ShapeLinker, a differentiated approach to de novo design of linker domains that utilizes multi-parameter-optimization via reinforcement learning to steer the design efforts in the desired chemical space. The system leverages SMILES (Simplified Molecular Input Line Entry System) data, which is a widely used notation for representing chemical structures as text strings, to optimize for certain physicochemical properties as well as 2D and 3D requirements within the linker.
This work contributes meaningfully to our toolkit for therapeutic development as PROTACs remain a promising therapeutic agent for various diseases, including cancer, where the targeted degradation of disease-causing proteins can have significant therapeutic benefits. The ShapeLinker approach offers a more rational and efficient approach for PROTAC design and optimization.
Can large language models democratize access to dual-use biotechnology? [Soice et al., arXiv, 2023]
Why it matters: Democratization of synthetic biology and machine learning tools, in combination with contract manufacturing organizations, make it much easier for anyone to design potential bioweapons. According to the authors of the papers, LLMs make this even easier.In this qualitative study, students at MIT investigated whether LLM chatbots could be prompted to assist non-experts in causing a pandemic. Whilst the short paper is a worth a read, in summary the students were able to obtain suggestions for “four potential pandemic pathogens, explained how they can be generated from synthetic DNA using reverse genetics, supplied the names of DNA synthesis companies unlikely to screen orders, identified detailed protocols and how to troubleshoot them, and recommended that anyone lacking the skills to perform reverse genetics engage a core facility or contract research organization”.
The paper highlights two important critical issues:
Existing evaluation and training processes for LLMs, which rely heavily on RHLF, are easily jailbroken with specific prompts.
A combination of LLM-led suggestions and CMO cybersecurity breaches could enable non-experts to successfully manufacture LLM-suggested pathogenic sequences by also hacking CMOs and labeling the sequences as ‘safe’.
However, we believe that the limiting step towards increased use of bioweapons, or purposefully caused pandemics, is not the retrieval of information. Whilst LLMs create a better interface, Google Search and the dark web have all the necessary responses to obtain the same level of information. The authors' suggestions to remove ~1% of all PubMed publications from LLM training to eliminate the possibility of prompting chatbots to deliver the pandemic-causing suggestions is likely not going to reduce any of the risk. Bad actors already have access to this information, and phishing scams are not new. As we highlighted in our Biosecurity piece, it will take a full stack ecosystem to reduce biothreat risk significantly.
MS2Mol: A transformer model for illuminating dark chemical space from mass spectra [Butler et al., ChemRxiv, June 2023]
Why it matters: The identity and structure of the vast majority of metabolites, the intermediates and endproducts of cellular processes in our body, is still unknown to science. Platform biotech Enveda Bio published a new transformer-based method for predicting the structures of unknown metabolites based on mass spec data. Enveda Bio, a platform biotech that analyzes plants for natural compounds with therapeutic potential, published a preprint this week describing a new method for identifying novel metabolites on mass spec. Mass spec is the workhorse method for characterizing metabolites in biological samples. Briefly, individual molecules in a sample are separated using liquid or gas chromatography, and then individual molecules are ionized and measured for their total mass, generating an MS1 spectrum. These molecules are then fragmented, and the constituent parts re-measured generating an MS2 spectrum.
Existing methods for predicting the structure of unknown molecules rely on existing databases of known molecules. Enveda developed a new method called MS2Mol that translates mass spectra for unknown compounds into chemistry structures. The model uses an encoder/decoder framework called BART. A masked language model was trained on spectra from a set of molecules, and then asked to predict the missing peaks on spectra from new molecules. The model was 50% better at predicting the structure of unknown metabolites than alternative approaches that rely on reference databases. The ability to predict structure is highly valuable for drug discovery workflows, as drug developers can assess whether the molecule possess drug-like properties, is easily modifiable by medical chemists, and whether it possesses similar chemical motifs to known drugs.
What we listened to
Notable Deals
Eli Lilly to acquire DICE Therapeutics in $2.4 billion deal
Verve snags $60M from Lilly to tackle Lp(a) with one-time gene editing
Astellas delves deeper into protein degrading drugs with Cullgen deal
Novo and Moderna seal base-editing pacts with Life Edit
Attovia, a new California biotech, launches with plans to rethink how drugs bind their targets
In case you missed it
Bessemer and XPRIZE launched the Deep Tech 100, the definitive ranking of the world’s top 100 private deep technology companies featuring a number of next-generation bio platforms.
Good Meat and Upside Foods become first company to sell cultivated meat approved by FDA and USDA
What we liked on Twitter
Scientists and their posters. @Caroline_Bartma
A conversation between Andrew Ng and Geoff Hinton on AI risks. @AndrewYNg
Scientific writing workshop and office hours. @NikoMcCarty /
As machine vision improves, the internal representations of the models are less human brain-like. @DrewLinsley
Tracking performance of AI tools in answering scientific questions. @andrewwhite01
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone