BioByte 045: partnership strategy user-manual biotechs, IRA's first drug list, epigenetic drivers of glioma, in vivo perturb-seq, synthetic data for clinical AI
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. If you’d like to connect or collaborate, please shoot us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone. Happy decoding!

Quick rundown of what’s good in bio this week:
a user-manual for equity and partnership strategy for biotech startups
the list of initial drugs potentially subject to price setting by the IRA
how to capture value as a deep tech AI company
blood-based biomarkers for cognitive dysfunction after COVID hospitalization
in vivo CRISPR screening for dissecting gene circuits involved in the tumor immune microenvironment response to radiation
synthetic data for improving medical AI for skin lesion detection
epigenetic bases for gliomas, the most common form of brain cancer
What we read
Blogs
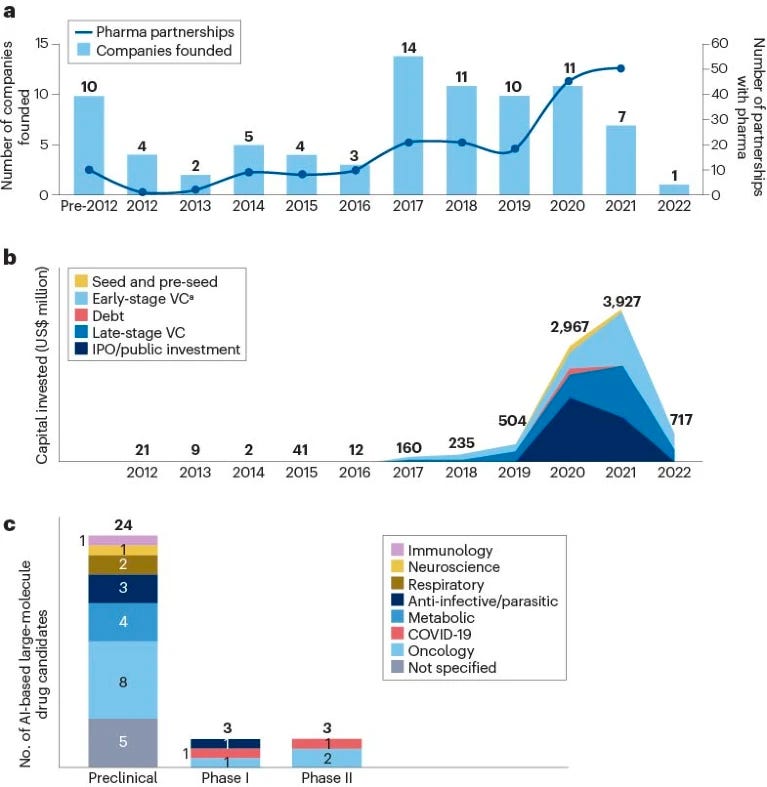
The company landscape for artificial intelligence in large-molecule drug discovery [Nagra et al., Nature Drug Discovery Reviews, 2023]
A succinct analysis of the AI-first biotech companies working on large-molecule discovery shows that:
Most (60%) we’re started within the last 5 years
“There is also some emerging evidence of consolidation amongst these companies, such as iBio’s acquisition of RubrYc Therapeutics in 2022.
The companies analyzed in this space raised US$3.9 billion in 2021, with $2.7 billion of this being raised by venture capital firms alone
Total investment declined significantly in 2022, however, to $0.7 billion.
Notable activity includes AbCellera and Absci raising $555 million and $200 million, respectively, in initial public offerings in 2020 and 2021, and Generate Biomedicines raising $370 million in series B funding in 2021.
Established large biopharma companies have also partnered with AI-driven biotech companies, with 51 partnerships identified in 2021, up from 10 partnerships in 2016”
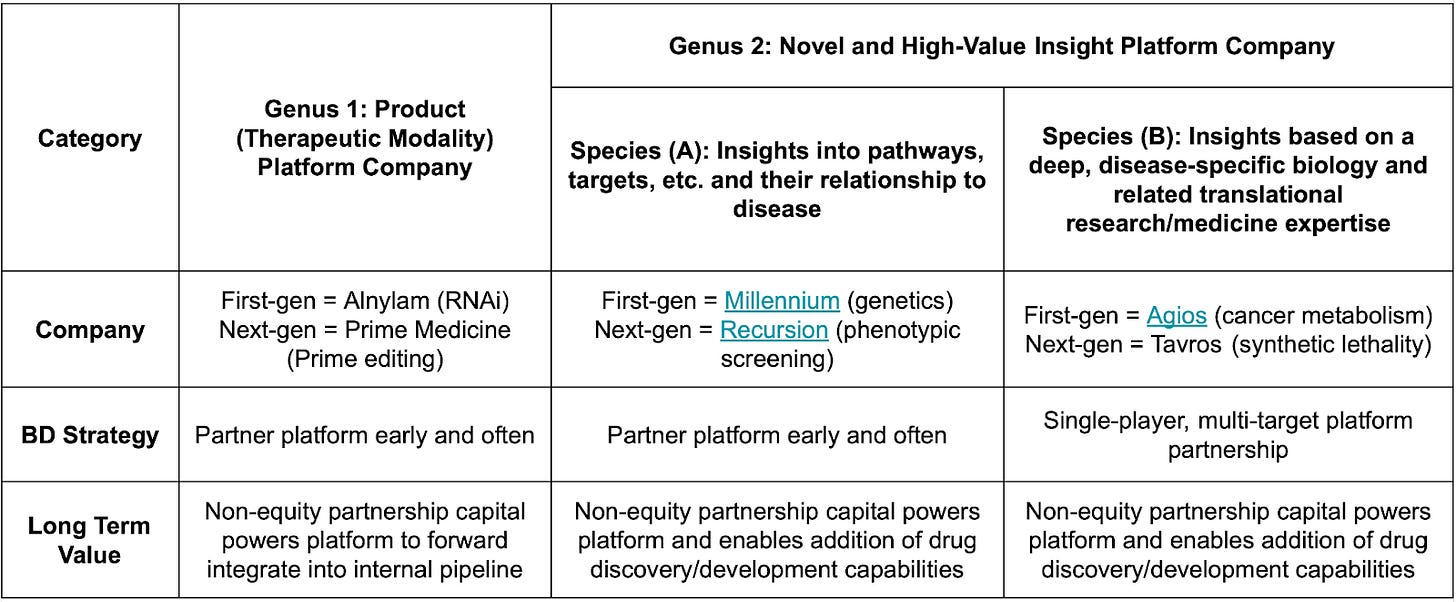
Early-Stage Biotech Value Creation: The Roles of Equity and Partnerships [Steve Holtzman, Aug 2018]
This blog from Steve Holtzman, former President of Decibel Therapeutics and Founder/CEO of Infinity Pharma, is 5 years old now, but is as relevant as ever. The essay discusses various fundraising and partnerships strategies for different types of biotech companies, including product companies that produce their own therapeutics and insight companies that deliver knowledge on targets, therapeutic MoA, etc to partners. Huge thanks to KdT Ventures fellow Emilio Ferrara for compiling the table below, which captures the key distinctions in BD strategy and value capture between different types of biotech companies.
Full-Stack Deep Tech Startups & AI Value Capture [Michael Dempsey, On My Mind, Sept 2023]
Mike Dempsey dives into the full-stack startup in deep tech and AI, which he defines as companies that build frontier technologies and deliver ROI themselves. Full-stack companies allow for long-term moat building and stronger value capture but don’t necessarily have the sexy point solution that makes other companies seem attractive. They are harder to build, involve many operational complexities, and often start pretty low margin. Mike sums up this concern well—
“all startups are hard and likely to fail, so you should build one with the largest scale of ambition and value capture that you believe in.”
He argues that deep tech companies create market cap destruction and expansion, which by nature gives rise to new business models and entirely novel ways of thinking about how things get done. One way to go about thinking about creating this type of business is how to leverage new technologies to consolidate and verticalize industries that inherently have inefficiencies. This is a particularly interesting thought experiment in bio where breakthrough technologies are coming out at an accelerated pace and understanding how to wield them to change the status quo over time can allow for massive full-stack opportunities. The challenge is managing the technology with organizational and operational complexities.
The Inflation Reduction Act’s First Drug List [Derek Lowe, In the Pipeline, 2023]
Derek Lowe provides his commentary on the first list of drugs that Medicare will be able to negotiate prices for as provisioned for by the Inflation Reduction Act. The takeaways:
List of drugs here
Medicare has, until now, been unable to negotiate drug prices
Most people want this, biopharma does not
The list starts at 10 drugs in 2026 and gradually increases to 60 by 2029
These drugs do not have market competition and cost Medicare a lot
Generic versions of the drugs may be available by the time price negotiation actually kicks in (in three years)
We might not actually get to price negotiation by 2026 because of all the legal battles the drug industry is launching on this
How much money this will actually save the healthcare system is TBD
Lawyers might be the real winners here ;)
Academic papers
Acute blood biomarker profiles predict cognitive deficits 6 and 12 months after COVID-19 hospitalization [Taquet et al., Nature Medicine, 2023]
One in eight patients receives their first ever neurological or psychiatric diagnosis within 6 months following COVID-19. These patients can suffer from cognitive deficits (brain fog), which are common, persistent and affect occupational functioning. The biological explanation to how cognitive deficits develop post-COVID-19 is unknown.
These cognitive deficits are complex, with both objective and subjective components, and are too heterogenous to assign a single biological mechanism. The authors of this study pose the hypothesis that different dimensions of cognitive deficits are predicted by different biological states. They used data from the large prospective longitudinal cohort study (PHOSP-COVID) to discover associations between biomarkers and post-COVID-19 cognitive deficits (measured 6 and 12 months later). The authors used canonical correlation analysis (CCA) as a dimensionality reduction technique, which enabled them to discover patterns of covariation between sets of variables.
Their analysis demonstrated that two dimensions link biomarkers with cognitive profiles. The first dimension linked high fibrinogen (relative to C-Reactive Protein (CRP)) to cognitive deficits after infection. The authors postulate this could be due to a combination of a hypercoagulable state and direct effects of fibrinogen on the brain (endothelial cell death). The second dimension linked high D-dimer (relative to CRP) to cognitive deficits and occupational impact after infection; partly mediated by shortness of breath and fatigue at 6 months. This is possibly due to associated coagulopathy causing brain microthrombi or pulmonary embolisms. Secondary analyses using large-scale EHR data also showed association with D-dimer is specific to COVID-19 but not with fibrinogen.
Thanks Ignacio Perez for the tip!
In vivo perturb-seq of cancer and immune cells dissects oncologic drivers and therapy response [Liu et al., BioRxiv, Aug 2023]
Why it matters: A new multiplex, single cell in vivo genetic perturbation method for dissecting gene circuits involved in the tumor immune microenvironment to inform our understanding of tumor response to radiation therapy. Pooled genetic perturbation screening is a high-throughput method that leverages gene editing enzymes like Cas9 to modify gene activity to study gene function. While in vitro genetic perturbation screens like Perturb-seq have advanced considerably, in vivo screens have been challenging. In this paper, in vivo perturb-seq was used to study the effect of radiation on glioblastoma (GBM) tumor cells. Guide RNA libraries, which are used to target and knock down activity of specific genes, were constructed to target genes associated with radiation sensitivity, resistance, and growth phenotypes. By comparing single cell RNA-seq of tumor cells both in vitro and in vivo in response to radiation therapy, the authors discovered that in vivo perturbation resulted in a more heterogeneous transcriptomic response to radiation (note the difference in heat maps showing gene expression changes for in vitro on the left and in vivo (CED) on the right in the figure below), highlighting the importance of perturbation screening in the proper physiological context. In vivo perturbation of macrophages was also performed to demonstrate the role of inflammation-related genes in driving cell-cell interactions within tumors. In summary, this approach for dissecting genetic circuits involved in the tumor immune microenvironment will be powerful for discovering and validating new genetic targets of cancer therapy.
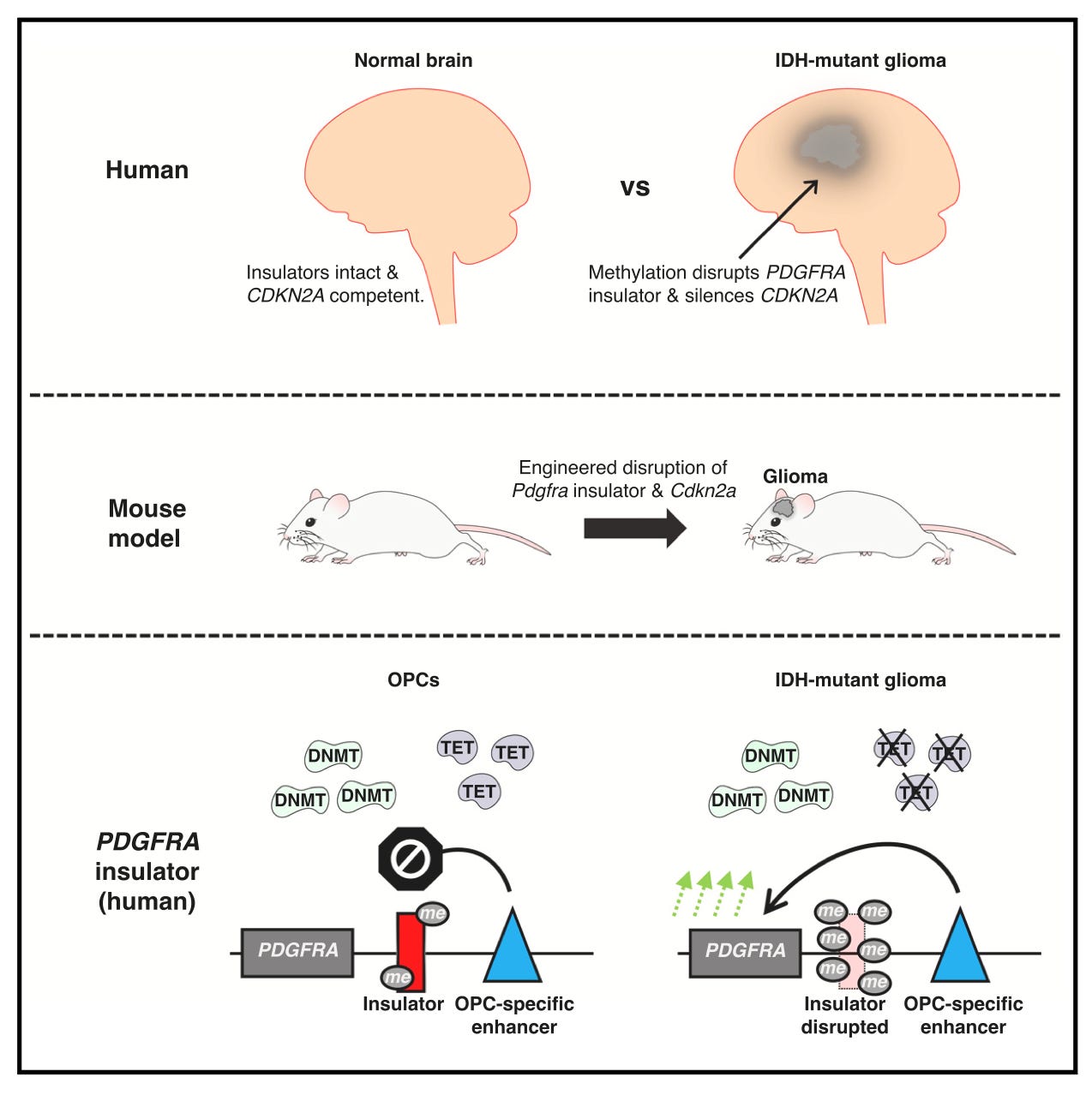
Modeling epigenetic lesions that cause gliomas [Rahme et al., Cell, 2023]
Why it matters: Gliomas are the most common form of central nervous system neoplasm and are notorious for their diffusely infiltrative nature that spread throughout the surrounding brain, making surgical and medical management challenging. These cancers are known to have a rich methylation profile, but such epigenetic variability makes it challenging to identify true oncogenic drivers vs. passengers. Here, the authors establish a new model for low-grade glioma (LGG) and identify PDGFRA insulator loss and Cdkn2a promoter silencing as drivers in isocitrate dehydrogenase (IDH) mutant gliomagenesis. LGGs with IDH mutations (associated with upregulated PDGFRA, contributing to oncogenesis) have significant DNA hypermethylation throughout their genome, which have been shown to silence promoters of tumor suppressor genes. The authors compared methylation status of CTCF insulators (regulatory elements that regulate transcription) near PDGFRA in IDH-mut and wild-type tumors and found that mutant tumors were associated with significant methylation of CTCF. Using CRISPR, the authors disrupted the CTCF insulator in mouse embryonic stem cell-derived neural progenitor cells and oligodendrocyte progenitor cells (OPCs), which have been identified as potential precursor cells for gliomas. They found that insulator loss allows an OPC-specific enhancer to activate PDGFRA and amplify OPC proliferation, while having no effect on NPCs which lack the enhancer. They additionally profiled several other methylation sites and found that CDKN2A is regularly methylated in IDH mutant gliomas, and found that this led to decreased P53 and increased OPC proliferation. The authors then perturbed Cdkn2a and CTCF in vivo and found that this led to hypercellularity, which was enhanced by PDGF supplementation (engaging PDGFRA), leading to malignant glioma. Thus, the authors successfully identify epigenetic drivers in IDH mutant gliomas.
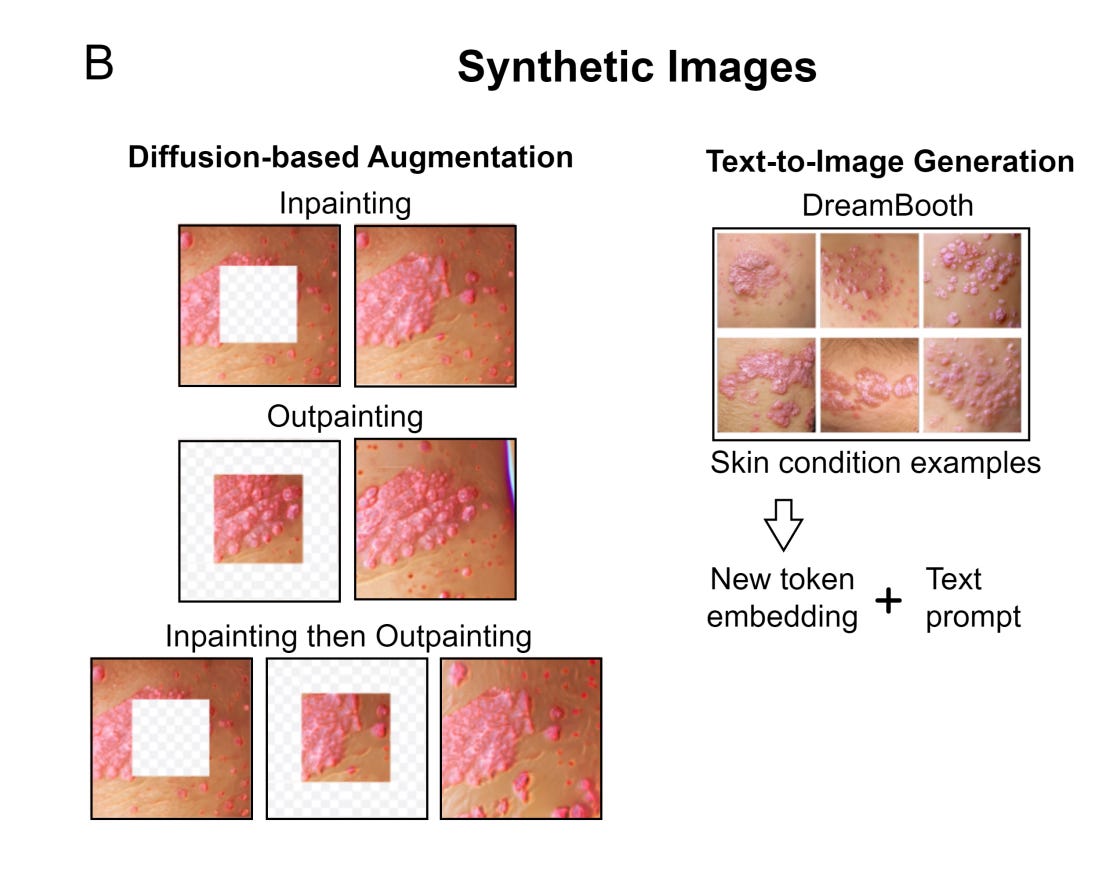
Augmenting Medical Image Classifiers with Synthetic Data from Latent Diffusion Models [Sagers et al., arXiv, Aug 2023]
Why it matters: This study explores whether synthetic medical images generated by AI can improve disease classifiers, especially for rare conditions or underrepresented groups. As large multi-modal diagnostic models mature in biomedicine, issues pertaining to data sparsity and missing data become more prevalent. This findings of this paper suggest that synthetic augmentation approaches are worth exploring further.In this study led by Dr. Raj Manrai, researchers used a state-of-the-art diffusion model to generate over 450,000 synthetic images across 9 skin conditions and a range of skin tones. They found that adding these synthetic images to the training dataset improved classifier performance, especially when real world data was limited. The benefits plateaued after a 10:1 ratio of synthetic to real images. Though the gains were smaller than simply adding more real data, synthetic images helped classifiers generalize.
Impressively, skin cancer detection improved across all skin tones when augmented with synthetic data. Overall, this research demonstrates synthetic data's potential to supplement real images for model development. Diverse real-world data remains most important, but synthetic data can provide a multiplier effect.
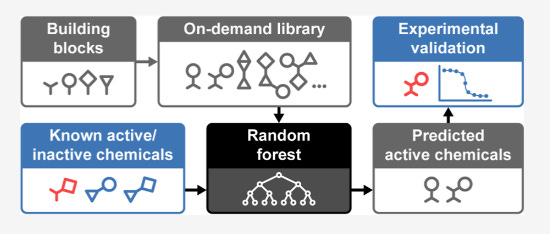
Evaluating Scalable Supervised Learning for Synthesize-on-Demand Chemical Libraries [Alnami et al., Journal of Chemical Information and Monitoring, Aug 2023]
Why it matters: This paper demonstrates a practical approach to leveraging machine learning models for virtual screening of massive chemical libraries. Selecting promising compounds from enormous synthesize-on-demand libraries could accelerate drug discovery.In this study, researchers developed a workflow to select a top performing random forest model for predicting bioactivity against a bacterial target. They prospectively tested this model on libraries of over 8 million commercially available compounds and over 1 billion synthesize-on-demand compounds. The model consistently achieved high hit rates, identifying bioactive and chemically diverse compounds. With only a small testing budget, it recovered potent hits from the massive synthesize-on-demand library that would likely be missed by simpler methods. The study shows supervised learning models can effectively prioritize virtual compounds for synthesis and guide experimental screening of gigantic chemical spaces.
What we listened to
Notable Deals
Novo, continuing deal streak, buys another obesity drug startup
Danaher to buy antibody supplier Abcam for $5.7B
Neumora, a richly funded brain drug developer, readies for an IPO
Former Google AI expert raises $100mn for biotech start-up
Nimbus tacks on $210M in funding as investors swat away IRA concerns
In case you missed it
STAT’s Healthcare Generative AI Tracker
What we liked on Twitter
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone









"all startups are hard and likely to fail, so you should build one with the largest scale of ambition and value capture that you believe in.” I love this quote. It call for founders and entrepreneurs to aim high and pursue ambitious goals, even in the face of obstacles. I am no founder but that quote goes hard 😆