BioByte 056: the first approved CRISPR medicine, LLMs for genetic discovery, generative models for protein complexes, super-additivity in combination therapies, new NVIDIA-Genentech collaboration
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. If you’d like to connect or collaborate, please shoot us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone. Happy decoding!
Here’s your weekly dose of tech x bio:
Vertex and CRISPR Therapeutics' Collaboration: The UK approved CASGEVY, the world's first CRISPR/Cas9 gene-edited therapy for sickle cell and beta thalassemia, developed through the collaboration between Vertex and CRISPR Therapeutics.
Med-PaLM 2 in Genetic Discovery: The use of the LLM Med-PaLM 2 in genetic research has accelerated genetic discovery, including identifying genes for various biomedical traits and developing a new model for hearing loss.
Combination Therapies in Cancer: Most FDA-approved combination therapies for advanced cancer are additive, not synergistic, suggesting a need for better translation of synergistic strategies from preclinical models to human trials.
Programmable Protein Design: Chroma, a generative model for protein design, represents a breakthrough in AI-driven drug discovery, enabling the sampling of novel protein structures and sequences.
Drug Distribution in Cellular Condensates: Research on biomolecular condensates highlights the importance of understanding chemical environments for effective drug distribution within cells.
Nvidia and Genentech Collaboration: Nvidia and Genentech's partnership leverages their respective strengths in ML and drug discovery to optimize ML-driven drug development processes.
What we read
Blogs
Vertex and CRISPR Therapeutics Announce Authorization of the First CRISPR/Cas9 Gene-Edited Therapy, CASGEVY, [Vertex], World’s first CRISPR medicine approved in UK for sickle cell, beta thalassemia [Ned Pagliarulo, Biopharma Dive, Nov 2023]
Casgevy (exagamglogene autotemcel), a CRISPR/Cas9 gene-edited therapy arising from the collaboration between Vertex and CRISPR Therapeutics, was approved in the UK to treat blood diseases sickle cell and beta thalassemia. This makes it the first gene edited therapy to be approved anywhere in the world.
Casgevy is an autologous, ex vivo CRISPR/Cas9 edited therapy. The patient’s own hematopoetic stem cells are edited to produce fetal hemoglobin in red blood cells. The biopharma companies estimate about 2000 people in the UK will be eligible which include “patients 12 years of age and older with SCD with recurrent vaso-occlusive crises (VOCs) or TDT, for whom a human leukocyte antigen (HLA) matched related hematopoietic stem cell donor is not available”.
Vertex splits all costs and profits of Casgevy 60/40 with CRISPR Therapeutics. Whilst the drug has been approved, commercialization is still hard. The price tag is expected to be in the millions and history shows that it will be a challenging market: Bluebird bio’s beta-thalassemia gene therapy was approved in 2019 and then withdrawn after it failed to secure enough reimbursement.
Betting on Deep Tech [Leo Polovets, Coding VC, November 2023]
Deep tech, including life sciences/biotech companies typically get a bad reputation for being capital intensive and having poor outcomes relative to time to exit. Leo Polovets, investor at Humba Ventures, uses this post to analyze Pitchbook data suggesting otherwise. He looks at deep tech returns and cites massive outcomes, many of which were in biotech (10X Genomics—$4.7B, Foundation Medicine—$5B) and hypothesizes more opportunities in the market for deep tech given how outdated the technologies of some industry incumbents remains. The argument that deep tech companies may be more capital intensive also seems flawed. After analyzing companies across both deep tech and traditional sectors, it appears the median funding raised to exit is 11-15% of its exit size. We see this risk/return profile in biotech where there is capital required to set up labs but meaningful exits are often much larger in scale accordingly. Perhaps most interesting, Leo actually found that deep tech companies are quicker to exit than traditional companies by about 10-25% with life science companies being the quickest. This could be due to IP moats built early, showing something of value for each company. Additionally, the odds of a good exit for a deep tech company might be higher but the number of such exits is lower, showing the relative scarcity and difficulty of building in deep tech today.
Academic papers
Genetic Discovery Enabled by A Large Language Model [Tu et al., BioxRiv, November 2023]
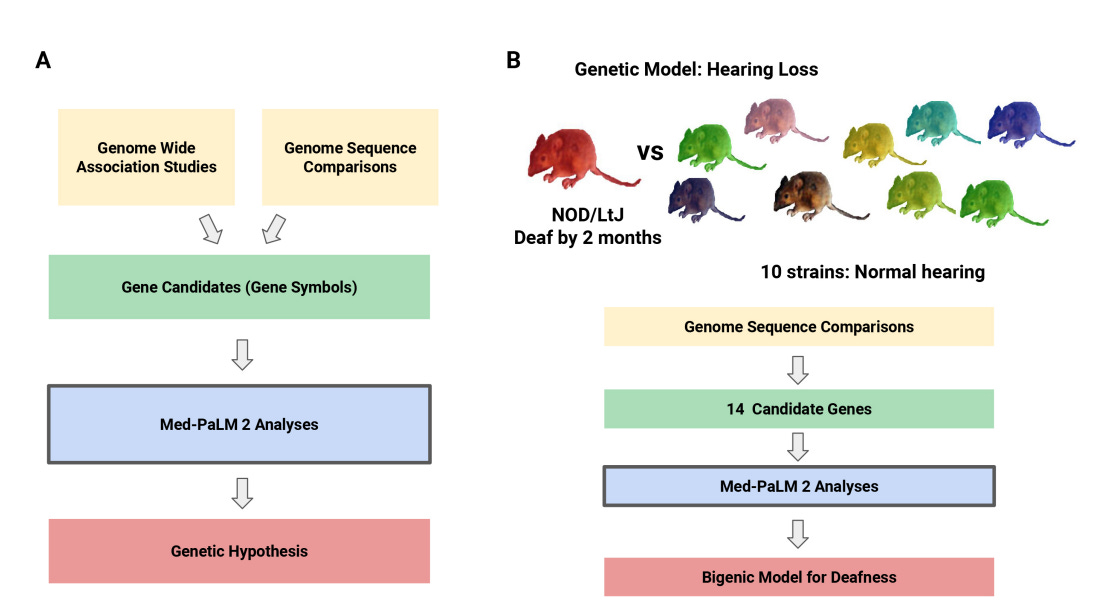
Why it matters: The use of LLMs like Med-PaLM 2 in genetic research represents a significant advancement in the field of biomedical science. This approach can accelerate the pace of genetic discovery by rapidly analyzing complex datasets and identifying key genetic factors linked to specific diseases or traits. The success of Med-PaLM 2 in identifying causative genes for various biomedical traits, including a novel genetic model for hearing loss, highlights the potential of AI-driven approaches in uncovering intricate gene-phenotype relationships.Tu et al. discuss the application of a specialized LLM, Med-PaLM 2, in genetic discovery, particularly in identifying causative genetic factors for various biomedical traits in mice. Med-PaLM 2 was tested for its ability to analyze sets of candidate genes from mouse genetic models and correctly identify genes responsible for six biomedical traits, including diabetes and cataracts. The model successfully identified the murine genes containing experimentally verified causative genetic factors for these traits. Furthermore, it was able to analyze a list of genes with high-impact alleles and identify a causative murine genetic factor for spontaneous hearing loss. This led to the development of a novel bigenic model for susceptibility to spontaneous hearing loss, demonstrating Med-PaLM 2's ability to generate novel hypotheses and facilitate genetic discovery.
Additivity predicts the efficacy of most approved combination therapies for advanced cancer [Hwangbo et al., Nature Cancer, November 2023]
Why it matters: Combination therapies are common in oncology to improve the response rate of monotherapies, and are often hypothesized to have a synergistic, or super-additive mechanism of action. A new analysis of FDA-approved combination therapies for cancer shows that the vast majority of combo regimens show additive, not synergistic effects.The field of oncology is increasingly shifting from monotherapy to combination therapies to improve efficacy and response rates, overcome drug resistance, and more effectively target tumor heterogeneity. Rational combination strategies seek to achieve a synergistic drug effect (response to A+B combo > A or B monotherapy) using knowledge of each drug’s mechanism of action. Despite a number of approved combination therapies, it is not clear whether these combinations have a synergistic (i.e., super-additive), additive, or less-than-additive effect.
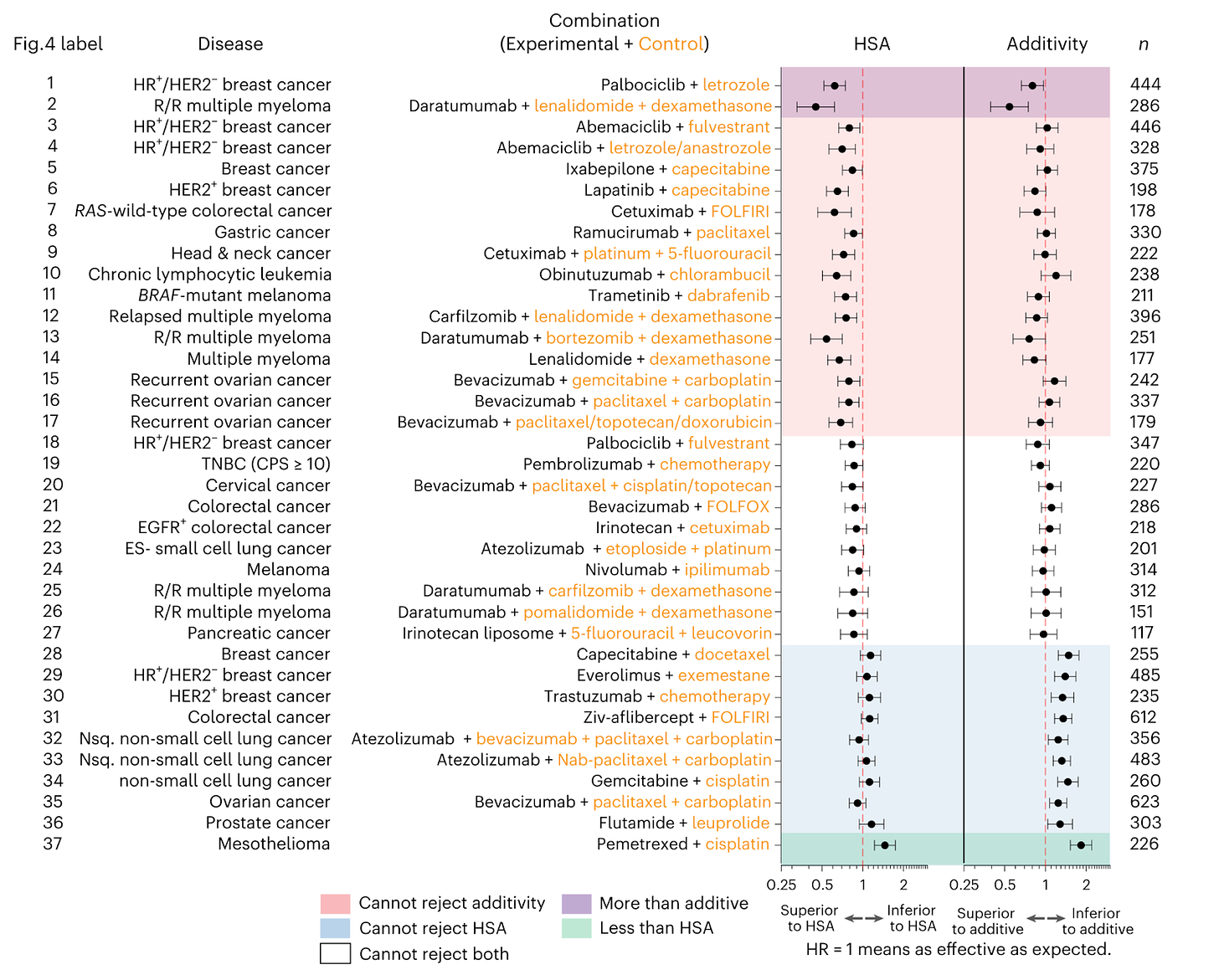
To tease apart these possible explanations for combo therapy efficacy in cancer trials, Adam Palmer’s lab at UNC developed a model of drug additivity for progression-free survival (PFS) to determine if drug combinations are additive or synergistic. All combination therapy trials in advanced cancers that led to FDA approval from 1995 to 2020 were analyzed. A model of additivity accurately matched clinical efficacy data for most approved drug combinations. Only 5% were synergistic or super-additive (purple in figure below). The additivity model could accurately predict clinical trial successes and failures, further demonstrating that most combination therapies work via an additive and not synergistic mechanism.
A big takeaway from this paper is that while combination therapies with additive effects are still superior to monotherapy and therefore clinically meaningful, we need better tools for translating synergistic strategies discovered in preclinical models into humans. One explanation for the inconsistency between preclinical and clinical evidence for drug combination synergy is tumor heterogeneity.
Illuminating protein space with a programmable generative model [Ingraham et al., Nature, November 2023]
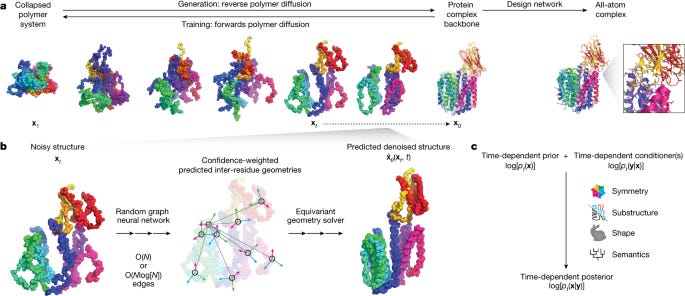
Why it matters: Protein molecules perform most of the biological functions necessary for life and can have novel applications across biology in health and agriculture. However, why be limited to the repertoire of known proteins? Scientists have desired to engineer custom proteins uniquely adept to serve a given function. The field of computational protein design aims to shorten this process by automating the design of functional proteins in a programmable manner. Chroma is a generative model for proteins and protein complexes that can directly directly sample novel protein structures and sequences and can be conditioned to steer the generative process towards desired properties and functions. It uses a combination of graph neural networks and diffusion. It is the third open-sourced de novo protein design engine after RF Diffusion and EvoDiff - an interesting trend for AI drug discovery companies. Generate Biomedicines (the creator of Chroma) believes drug discovery is more than proprietary models and the real value may lie in differentiated data generation capabilities to feed the model. Generate has invested around $40M into its structural biology labs.
Distinct chemical environments in biomolecular condensates [Kilgore et al., Nature Chemical Biology, September 2023]
Why it matters: A better understanding of condensate biochemistry is critical to control where drug molecules distribute within a cell. In drug discovery, ideally all of the administered molecule should home to the target and not get ‘stuck’ in other cellular compartments. If this happens the dose administered will need to be increase which can increase the chance for off-target effects / toxicity.This paper investigates the prediction of where a given molecule will accumulate, using machine learning. The researchers devised a representative system to study what might be causing drugs to concentrate in one condensate over another. First the researchers assembled 3 types of artificial condensates: gene transcription, regulation and the nucleolus. They then isolated the dominant protein that forms the framework for each of these three types and formed simplified versions of the condensates just containing the protein. They formed a diverse library of 1500 small molecules and tested how strongly each molecule would concentrate in each condensate. Finally an ML model was trained on the resulting data. The team found that molecules which favor a specific type of condensate tend to have shared chemical features.
Notable Deals
GLP-1s - one of the most lucrative drug classes in history?
Current global sales of GLP-1s are estimated at around $35B in 2023 and projected to reach $103B by 2030 with $41B project from obesity. The obesity space has also seen strong clinical data in Novo’s (SELECT) CV outcomes. Lilly and Novo are making large investments in manufacturing capacity (Lilly expanded $2.5B facility in Germany) and are poised to dominate the market. However there are 95+ weight loss agents being developed by 42+ companies in novel combinations and modalities to address every type of consumer and potential side effect.
Nvidia / Genentech - the quintessential compute company and biotech join forces
Overview: Nvidia and Genentech are collaborating to use their respective expertise to further ML-driven drug discovery.
Nvidia will help Genentech optimize its ML deployment by accelerating training and inference of Genentech’s 20 drug discovery models. Genentech is pioneering a ‘lab-in-a-loop’ where experimental data is fed into ML models which are improved, deploy results for testing and these are re-fed into the cycle. It is time-consuming for these models to adjust to new data and Nvidia will help improve their efficiency.
Genentech will deploy their models on Nvidia’s DGX cloud and will help Nvidia gain in insights into AI-related challenges in drug discovery and development. It will use these learnings to improve its BioNeMo platform and cement itself in the deep learning age of drug discovery.
Quotient raises $50M to spot evolutionarily adept cells with somatic genetics
Flagship’s latest launch is its first foray into the UK and leans on crucial work from scientists at the Wellcome Sanger institute.
Mission: identify somatic cells that have either acquired susceptibility or resistance to disease overtime and harness these traits for therapeutic development.
Therapeutic area: The company claims its approach is applicable to all indications and has not indicated a potential focus.
As one Flagship-built biotech launches, another fails
Evelo Biosciences will dissolve after finding no viable path to a new funding and research strategy. Evelo’s platform was based on novel monoclonal microbials targeted against inflammatory conditions and produced EDP2939 that was tested in Phase II for Psoriasis but ultimately didn’t meet its primary endpoints. The company experienced recent clinical trial failures in atopic dermatitis and considered partnered parts of its platform but still found no path to continue. This highlights the difficulty of novel biology in drug discovery, especially the complexity of microbiome-based therapies which are a relatively new and complex field. Evelo had prominent investors and backers such as GV, Alexandria Venture Investments, Celgene and longtime Biogen chair Stelios Papadopoulos.
What we listened to
Atlas Venture 2023 Year in Review - Bruce Booth
In case you missed it
Scaling Biology: Chris Gibson, Co-Founder and CEO of Recursion Pharmaceuticals [Pablo Lubroth, Decoding Bio]
What we liked on social
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone