BioByte 064: LLMs for neuro, bias in protein language models, engineered T-cell therapeutics inspired by T-cell cancers, AI for molecular glues
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. Happy decoding!
Happy Valentine’s Day fam!
Quick housekeeping before we dive in: we’re launching a podcast. Let us know who you want to hear in conversation - DMs (twitter/LI) are open.
What we read
Blogs
Iambic Therapeutics Announces New Research Published in Nature Machine Intelligence Demonstrating the Capabilities of its Generative AI NeuralPLexer Technology to Predict Protein-Ligand Complex Structures [Iambic Therapeutics, Feb 2024]
And so the protein-ligand structure prediction wars continue… last year we saw posts from David Baker’s group, Google’s Isomorphic Labs and now we see Iambic Therapeutics follow with comparison results for their NeuralPLexer model. The comparison centers around a benchmark called posebusters, initially chosen by the Baker group and adopted by the others for a ‘like-for-like’ comparison. The results Iambic post show slightly worse performance than Isomorphic’s AlphaFold-latest, but the team also report on a subset where the model performs with high confidence. The team report on full complex prediction with no site specification and with site-specific specification, the latter being a much easier task. But how do these benchmarks actually translate into real drug discovery impact? That remains to be seen.
Academic papers
Large language models improve annotation of prokaryotic viral proteins [Flamholz, Nature Microbiology, Jan 2023]
Why it matters: Poor annotation of viral genomes in metagenomic samples is a hurdle to understanding viral diversity and function. The authors’ protein language model expanded the annotated fraction of a dataset by 29%. Expanding annotation of remote virus families can have an impact in the identification of useful genomic sequences, such as integrases and new capsids useful in gene editing and delivery.Current annotation approaches rely on alignment-based sequence homology methods. These are limited by lack of characterized viral proteins, wide divergence among viral sequences and lack of universal conserved marker genes to enable broad taxonomic analysis.
The authors used a protein language model (PLM) to improve upon homology-based methods. PLMs can capture physico-chemical properties of amino acids and resolve protein structure and function information from sequence. Viral proteins conserve structure and function more than sequence due to biochemical and fitness constraints. So using a PLM to classify viral protein families based on function rather than sequence, should improve viral protein annotation.
Using its PLM-based classifier to expand annotations on a database of ocean virome sequences led to a 29% improvement over homology-based methods.
Variant in the synaptonemal complex protein SYCE2 associates with pregnancy loss through effect on recombination [Steinthorsdottir et al., Nature Natural and Structural Biology, Jan 2024]
Why it Matters: Understanding the genetic underpinnings of pregnancy loss is crucial for developing targeted interventions and counseling strategies for affected individuals. This research identifies a specific genetic variant associated with pregnancy loss, offering potential markers for assessing risk and guiding research into therapeutic approaches. By elucidating the role of the synaptonemal complex and recombination processes in fertility, this study lays the groundwork for advances in reproductive medicine and genetic counseling, with the ultimate goal of reducing the incidence of pregnancy loss.The study uncovers a significant association between a variant in the synaptonemal complex protein SYCE2 and increased risk of pregnancy loss, through a detailed genome-wide association analysis involving over 114,000 women. This variant, SYCE2:p.His89Tyr, also correlates with specific recombination phenotypes, such as the more random placement of crossovers and altered recombination rates across chromosomes, emphasizing the role of recombination quality in pregnancy outcomes. This work represents the largest dataset investigation of its kind to date, providing new insights into genetic factors contributing to pregnancy loss.
Data science opportunities of large language models for neuroscience and biomedicine [Bzdok et al., Cell, February 2024]
Why it matters: This article outlines the expansive potential of LLMs within the realms of neuroscience and biomedicine, emphasizing how large-scale language models can help scientists re-evaluate classical neuroscience questions, enhance neuroscience datasets with advanced text sentiment, summarize vast amounts of information to bridge community divides, and integrate disparate information sources relevant to brain research. LLMs are poised to transform the landscape of neuroscience and biomedicine by facilitating a deeper understanding of cognitive concepts and phenomena in the brain.The article explores how LLMs might revolutionize our approach to neuroscience research and biomedical applications – unlocking new insights into the human brain and disease mechanisms, leading to improved diagnostic tools, treatment strategies, and a deeper understanding of the complex interplay between biological processes and cognitive functions.
The review discussed a mega-analysis of over 6,500 text reports of hallucinogenic experiences induced by 27 different drug compounds, which was integrated with drug-specific receptor binding profiles of 40 neurotransmitter subsystems in the human brain. This study utilized NLP algorithms to analyze free-form text reports from drug users, identifying coherent semantic themes and cross-modal synthesis patterns that shed new light on the mechanistic basis of psychedelic drug experiences and their impact on conscious awareness.
Addressing the Antibody Germlines Bias and Its Effect on Language Models for Improved Antibody Design [Olsen et al., BioRxiv, Feb 2024]
Why it matters: A new preprint from the Oxford Protein Informatics Group highlight an important source of bias in public training data of protein language models for antibody design in which antibodies that have undergone affinity maturation are under-represented.Protein language models (PLMs) have advanced considerably in recent years. PLMs are often pre-trained on large public databases of protein sequences such as the Observed Antibody Space for antibody-specific PLMs. A new preprint from the Oxford Protein Informatics Group describes an important source of bias in these data that affect the performance of PLMs for antibody design.
An antibodies specificity for an antigen arises from a region known as the complementarity-determining region (CDR). Antibodies acquire the ability to bind different antigens through several mechanisms. Before infection with a pathogen, a process called V(D)J recombination randomly assembles variable (V), diversity (D), and joining (J) gene segments in the immunoglobulin genes during B-cell development. After infection, somatic hypermutation introduces point mutations into the V(D)J region after antigen exposure, further diversifying the antibody repertoire. Antibody sequences generated through somatic hypermutation are referred to as non-germline because they are not inherited.
The problem for PLMs is that most the training data for antibody language models comes from blood samples, which has a lower proportion of affinity-matured antibody producing B-cells that have undergone somatic hypermutation vs other tissue samples such as from lymph nodes. In short, the highest-affinity, most specific antibodies are under-represented in the training data, and naive B-cells are over-represented (image below).
The authors found a germline bias within the OAS database, where the majority of B-cells have not undergone affinity maturation. This bias translated to the most popular PLMs, including Meta's ESM-2 and antibody-specific PLMs like AntiBERTy. To reduce bias, new models were trained optimized for predicting non-germline residues. The new models resulted in greater sequence diversity of generated antibodies, the greater binding affinity was not tested experimentally.
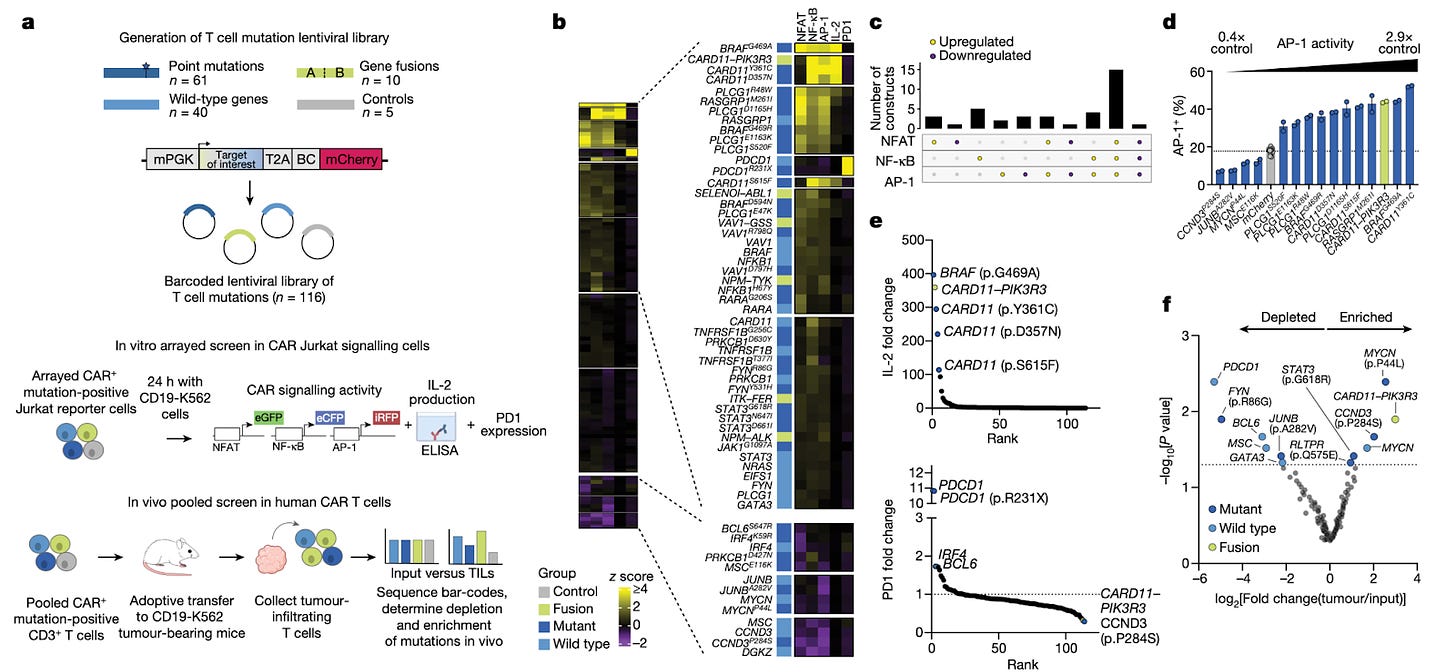
Naturally occurring T cell mutations enhance engineered T cell therapies [Garcia et al., Nature, 2024]
Why it matters: T-cell therapies have quickly become another pillar of the therapeutic armament, but need significant engineering to overcome poor in vivo persistence and modest efficacy in solid tumors. In this paper, the authors draw inspiration from T-cell neoplasms and their ability to survive in complex environments, and use this to improve cell therapy. They probe the landscape of T-cell cancers and identify mutations – particularly, a fusion protein of CARD11-PIK3R3 – that, when introduced into engineered T-cells, significantly bolster tumor cytotoxicity. This work further demonstrates how profiling cancer can yield insights that significantly improve therapeutic engineering. T-cell neoplasms are marked by a mutational landscape that significantly increase their survival in immunosuppressive environments. The authors identified 71 mutations of clonal T-cell neoplasms and cloned these genes into a library before transducing them into CAR-T or TCR-T cells. They found that introducing the fusion CARD11-PIK3R3 (identified in CD4+ T-cell lymphoma) into CD8+ therapeutic T-cells resulted in significantly improved survival in tumor-bearing mice. Additionally, they found that introduction of this fusion gene did not demonstrate any evidence of malignant transformation for up to 418 days after T-cell transfer. Of note, these CARD11-PIK3R3-bearing T-cells were extremely potent; about 20,000 of these cells were able to treat lesions extensively (as compared to traditionally needing ~1 million cells).
Notable Deals
Acquisitions
Gilead to buy liver disease biotech Cymabay for $4.3B - the acquisition will hand Gilead an experimental drug for primary biliary cholangitis. The FDA has just accepted the drug application and an approval decision will be made around August 2024.
Partnerships
VantAI and Bristol Myers Squibb partner to use geometric deep learning for the rational design of novel molecular glues - the deal involves undisclosed targets of interests and has a total value of $674M. Given that this figure is not broken down into upfront and milestones, it is difficult to delineate the near-term cash that this deal provides.
Financings
BioAge raises $170M on the obesity hype - Its lead candidate, azelaprag, is meant to help optimize weight loss from GLP-1 drugs so that muscle mass is preserved. The drug mimics the hormone apelin which is secreted in response to exercise. Although one of many programs for the anti-aging biotech, due to the obesity hype it has sparked investor interest from Lilly, Sofinnova Investments and Amgen.
Alys, a biotech based in Boston and Geneva, has launched with $100M funding - created through the merger of six Medicxi portfolio companies. Each company, now a subsidiary under Alys, targets a specific immune-related skin disease. This structure diversifies Alys's asset portfolio, reducing investment risk compared to single-asset approaches. The funding will support the subsidiaries in starting human trials for their medicines by the end of 2026.
Investors pile into psychedelic start-ups tackling mental health - “It is the second-highest month of fundraising ever recorded, after March 2021. Multiple people who work in the sector said they expected more backing for psychedelics groups, as promising scientific data and positive signals from regulators attract high-profile investors.”
The UK’s Science and Technology Secretary announces £100M Engineering Biology investment - the funds are aimed at research that bolsters pandemic preparedness, revolutionizing farming practices, and fortifying against floods.
BigBiome raises £310k in pre-seed investment to develop sustainable pesticides
What we listened to
In case you missed it
What we liked on socials channels
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone