BioByte 066: CAR-T for MS, will AI replace animal testing, diagnosis of ultra-rare monogenic disorders, BigRNA: a foundation model for RNA biology, Matter Neuro raises $26M to “cure unhappiness"
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. Happy decoding!
Hello team!
Last summer we hosted our inaugural AI x Bio Summit at the NYSE. We’re planning the 2024 edition which will likely be in the Bay Area. If you have any ideas for the programme, would like to speak at the event or attend it do message us (twitter/linkedin/email open).
What we read
Blogs
CAR-T therapy for multiple sclerosis enters US trials for first time [Mullard, Nature News, February 2024]
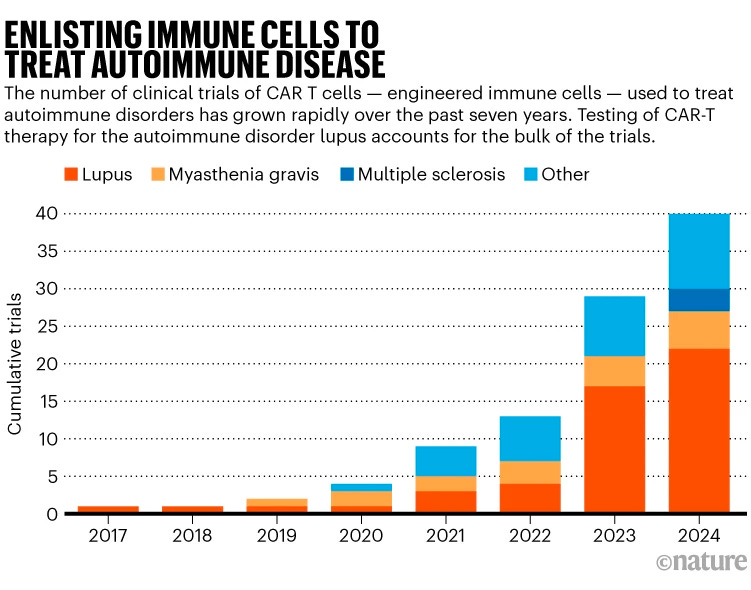
The initiation of the first U.S. trials for CAR-T therapy in treating multiple sclerosis (MS) marks a pivotal moment in the search for innovative treatments for autoimmune diseases. Renowned for its success in battling certain blood cancers, CAR-T therapy employs engineered T cells to target and eliminate problematic cells. In the case of MS, a disease characterized by the immune system's attack on nerve cells, this approach offers a beacon of hope. The trials, led by Kyverna Therapeutics and in partnership with researchers at Stanford University and Bristol Myers Squibb, aim to explore the potential of CAR-T cells in not just slowing but halting the progression of MS by targeting B cells implicated in the disease's pathology.
The significance of these trials extends far beyond the realm of MS treatment, embodying a broader quest to harness the immune system's power against a variety of autoimmune disorders. If successful, CAR-T therapy could represent a "one-and-done" approach, dramatically shifting the treatment paradigm for autoimmune diseases. This potential to reset the immune system safely and effectively could transform the lives of millions suffering from autoimmune conditions, offering a more straightforward, potentially less risky alternative to existing therapies such as autologous haematopoietic stem-cell transplantation. As the scientific community keenly observes these trials, the success of CAR-T therapy in MS could pave the way for a new era in the treatment of autoimmune diseases, underscoring the importance of innovative, targeted therapies in advancing patient care and outcomes.
When can AI deliver the drug discovery hits? [Asher Mullard, Nature Reviews Drug Discovery, Feb 2024]
The results from the Critical Assessment of Computational Hit-finding Experiments (CACHE) competition were recently published. The CACHE competition is modeled off of the CASP protein structure prediction competition, which rose to prominence with AlphaFold several years ago. The goal of the CACHE competition is to help benchmark molecular design methods by providing unbiased, high quality experimental feedback on computational hit-finding predictions. Twenty-three teams competed to predict which ligands would bind to LRRK2’s WDR domain (a previously undrugged pocket of a target relevant for Parkinson’s disease). Entrants could use whatever computational tools and methods they wished to design small molecule binders, which were then synthesized and tested in the lab to confirm activity.
There were a number of surprising observations based on the results:
In aggregate, all of the teams generated predictions for 2000+ potential binders, yet only 12 successfully bound the WDR pocket when tested in the lab, resulting in a <1% hit rate. This could be due to the limitations in the methods, but also due to difficulties in drugging the WDR domain itself.
Pat Walters, Chief Data Officer at Relay Therapeutics summed up the results well: “We're making incremental changes and improvements, but it wasn't like there was anything earth-shattering that came out of this first challenge.”
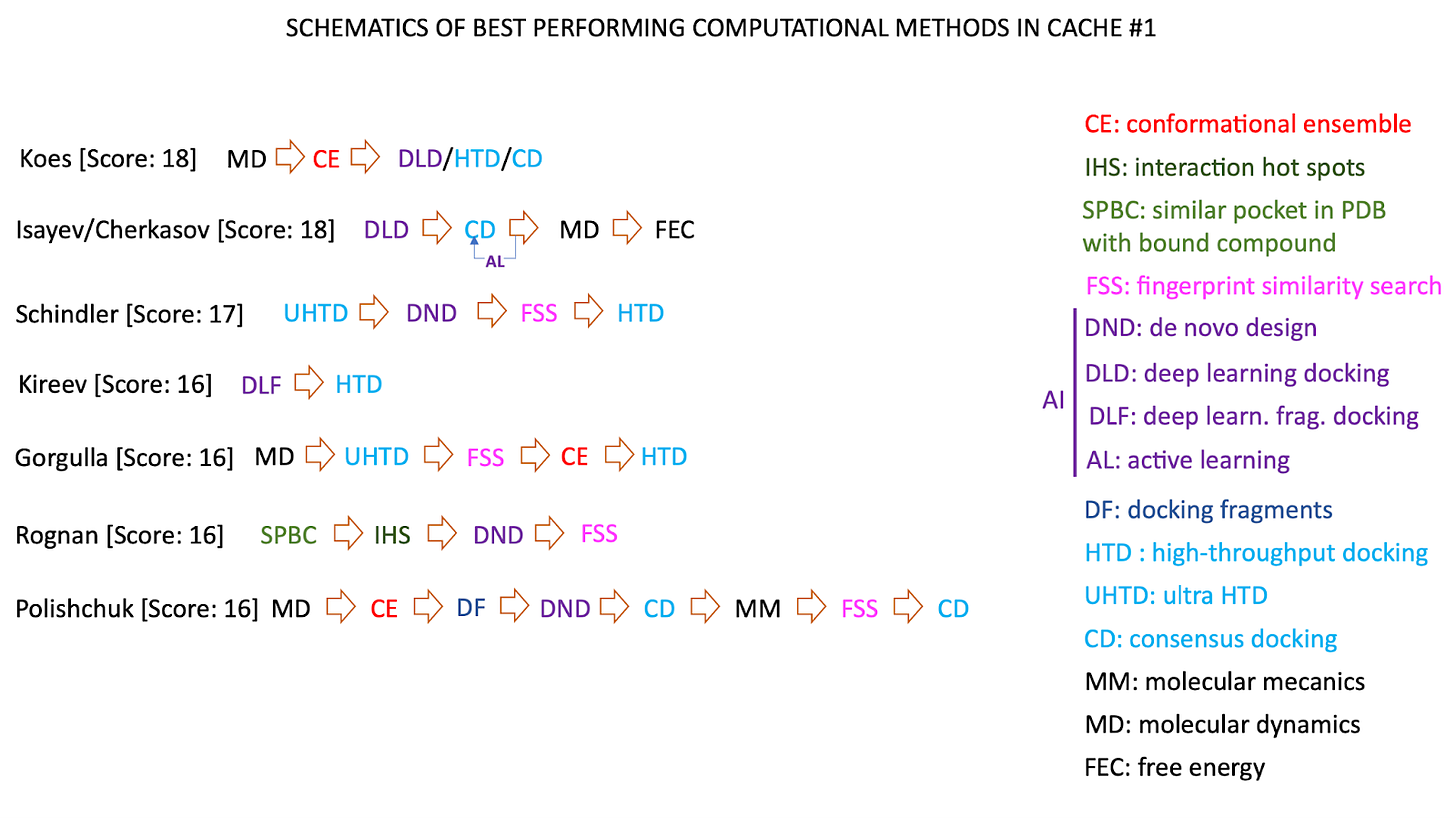
There weren’t any consistent patterns across types of methods used by the winning teams. Some teams used mostly machine-learning based methods, others used strictly physics-based methods with little/no ML. The figure below captures the complex variation in methods used across the various teams. The take-home is that there is unlikely to be one method/algorithm to rule them all, and state-of-the-art performance is likely to be highly context-dependent based on the target, binding pocket, etc.
Future iterations of the CACHE competition will explore additional targets, including three competitions currently ongoing for NSP13 helicase, the NSP3 macrodomain of SARS-CoV-2, and CBLB, an E3 ligase relevant for the design of protein degraders.
How AI will Replace Animal Testing [AI + Bio + Design, Brandon White, February 2024]
Animal testing is no stranger to scrutiny by biomedical researchers and the general public alike. Despite being the gold standard for research, animal trials are costly, time-consuming, and their efficacy and predictive validity hotly contested—why do 90% of drugs fail even after passing animal tests? Brandon White takes to his latest blog post to argue that AI can replace much of animal testing, a total paradigm shift. Brandon points out that with the FDA Modernization Act, there are policy pressures in place to “replace, reduce, and refine” animal testing. No longer does a drug absolutely have to undergo animal testing to be approved, although that is still the status quo. Companies outside of biotech like those focused on cosmetics are also adopting new practices including artificial skin models and bans on animal testing cosmetic products. Even big biopharmaceuticals like Roche and Eli Lilly have made commitments to reducing reliance on animal models, though alternatives are not immediately clear. Brandon suggests that multi-modal AI research can tie together various data sources to create models of biology. Though in the early stages today, he is confident we will be able to use such foundation models to predict efficacy & toxicity, help run trials, and troubleshoot experiments. In the short-run, focus is on building those robust datasets.
Academic papers
Joint, multifaceted genomic analysis enables diagnosis of diverse, ultra-rare monogenic presentations [Kobren, bioRxiv, February 2024]
Why it matters: This research underscores the power of integrating genomic data analysis across individuals with rare diseases to improve diagnosis and understanding of genetic disorders. By employing sophisticated statistical tools and a comprehensive evaluation framework, the study exemplifies how collaborative, data-driven approaches can enhance the identification of genetic causes for undiagnosed diseases. This breakthrough has significant implications for personalized medicine, offering hope for patients on prolonged diagnostic odysseys and paving the way for targeted therapeutic interventions.
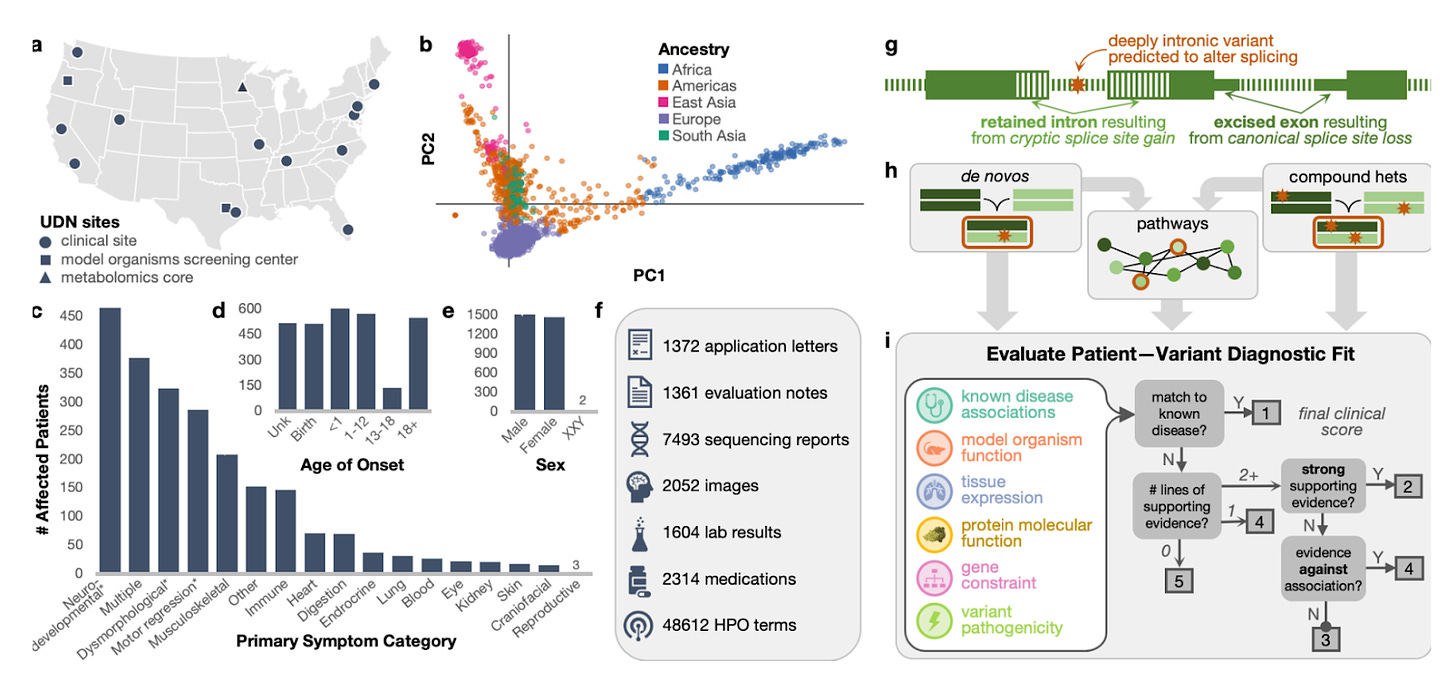
The study presents a comprehensive genomic analysis within the Undiagnosed Diseases Network (UDN), focusing on individuals with ultra-rare monogenic disorders. It introduces advanced statistical methods for analyzing whole genome sequencing data, identifying disease genes through de novo mutations, compound heterozygosity, and enriched pathways. This multifaceted approach not only confirmed known diagnoses but also unveiled novel disease-gene associations. The analysis highlights the significance of joint genomic analyses across diverse cohorts, showing that combining individual rare case analyses (N-of-1) with broader cohort-wide investigations can reveal new insights into genetic diseases.
An RNA foundation model enables discovery of disease mechanisms and candidate therapeutics [Celaj et al., 2023]
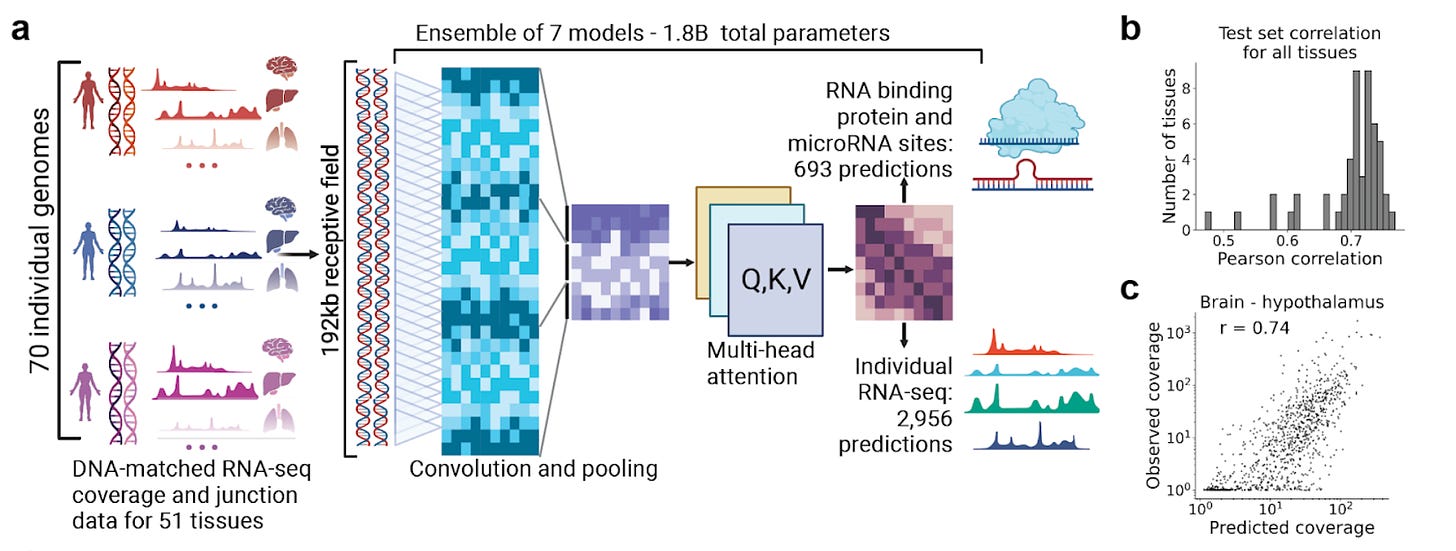
Why it matters: predicting the impact of transcriptional perturbations on regulatory networks has been challenging. The authors built a foundation model based on paired genotype and high resolution RNA expression data to carry out predictive tasks such as RNA binding protein specificity and microRNA binding sites; as well as finding oligonucleotides which could advance the field towards personalized therapeutics.
Last fall, the team at Deep Genomics published BigRNA: a foundation model for RNA biology that can be used in a range of tasks including prediction of tissue-specific RNA expression, splicing, microRNA sites, and RNA binding protein specificity from DNA sequence. The model was also used to identify pathogenic non-coding variants. Importantly, the model was used to predict the effects of steric blocking oligonucleotides (SBOs) that affected splicing and also used to design SBOs that blocked inhibitory regions to increase the expression of a disease-associated gene. BigRNA improved significantly over specialized models like TargetScan, DeepRiPe, SpliceAI and Enformer.
Notable Deals
Financings:
Curve Therapeutics raises $51.4M Series A to advance its small cyclic peptide-based discovery platform and its two preclinical oncology programmes into Phase I. Curve’s targets are HIFs (which regulate many genes that promote blood vessel growth in tumors) and ATIC (which is shown to be upregulated in cancers such as melanoma). The financing was led by Pfizer Ventures.
Matter Neuroscience raises $26M to “cure unhappiness” - the round saw participation from Polaris Partners and ARCH. Matter is creating an app to help users understand their patterns of thinking and how this affects them on a molecular level.
ORI Capital closes a $260M second fund, riding on the success of CG Oncology. The fund plans to invest across cancer, metabolic disease and neuro.
Partnerships:
Novo Nordisk and Neomorph partner to develop molecular glues - the promising technology aims to enabling the targeting of many proteins that may not be amenable to small molecule drug discovery. The total deal value is up to $1.4B, although the upfront is undisclosed. Novo hopes to use the technology to tackle difficult-to-drug targets in metabolic diseases, part of its strategy to back-fill its pipeline with innovative technologies.
What we listened to
AI Frontiers with James Zou: The Future of Multi-Modal AI in Medicine
In case you missed it
Fresh from the biotech pipeline: record-breaking FDA approvals [Melanie Senior, Nature Biotech, 2024]
What we liked on socials channels
Events
MIT Grand Hack 2024 – MIT Hacking Medicine [April 19-21]
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone