BioByte 068: Cell therapy for solid tumors, using a monoclonal for allergy, improving AlphaFold2 performance, using ML to understand small RNA secretion, Insilico's development of a TNIK inhibitor
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. Happy decoding!
Welcome to Spring. If there’s one thing we’ve learned, it’s that seasons are ephemeral, but Decoding is forever. Almost like that Hotel California line: “You can check out any time you like, but you can never leave!”
What we read
Blogs
First cell therapy for solid tumors heads to the clinic: what it means for cancer treatment [Sara Reardon, Nature, 2024]
Thirty-five years ago, Rosenberg et al. (1998), demonstrated the use of tumor-infiltrating lymphocytes (TILs) for the treatment of patients with metastatic melanoma. Now, this treatment, Lifileucel (produced by Iovance), is finally approved in the US, and represents the first cell therapy approved for treating solid tumors. Although we’ve seen significant success in using CAR-T therapies to treat cancer, these have only been well established in hematologic malignancies, such as leukemia and lymphoma. The approval of Lifleucel sets the foundation for establishing cell therapy as a promising modality for solid tumors.
Simply put, TILs are white blood cells (think along the lines of CD8+ cytotoxic T-cells) that have been found to penetrate solid tumors and destroy cancer cells. Superficially, the process is simple: surgeons harvest a patient’s tumor and send them to a lab, where they isolate TILs before expanding them. Patients are then treated with IL-2, which helps support TIL engraftment. In a trial testing Lifileucel in melanoma, 31% of participants had a reduction in tumor size; in a subsequent trial, 20% of melanoma patients receiving TIL therapy went into complete remission.
It’s important to note that TILs are bespoke for each patient – responding to specific antigens that aren’t known – and embodies truly personalized medicine. Given that each patient’s TILs may respond to an entirely different set of antigens, the FDA approved Iovance’s processes for cell expansion and administration to cancer patients. Although expensive ($515,000), several academic labs and companies are developing novel ways to improve the TIL manufacturing process, and are also applying them to other tumors. This work is exciting as it sets the stage for ushering in a new era of cell therapy.
New England Journal of Medicine Publishes Phase III Data on Xolair [Genentech, February 2024]
The New England Journal of Medicine published Phase III data from the OUtMATCH study, an NIH-sponsored study showing that Xolair (omalizumab) significantly reduced allergic reactions across multiple foods in people with food allergies. The study demonstrated that treatment with Xolair increased the amount of peanuts, milk, egg, wheat, and tree nuts that participants could consume without experiencing moderate to severe allergic reactions. Xolair was recently approved for children and adults with IgE-mediated food allergies, the first and only medicine on the market for that purpose. It has also been used to treat asthma and chronic urticaria, the first indications the drug was pointed towards. The safety findings were consistent with Xolair's known safety profile across its approved indications and in previous clinical trials.
In the study, 180 patients aged 1 to 55 years who were allergic to peanuts and at least two other common foods were treated with Xolair or placebo for 16 to 20 weeks. The results showed a statistically significant higher proportion of patients receiving Xolair could consume at least 600 mg of peanut protein and at least 1,000 mg of milk, egg, and cashew protein without experiencing moderate to severe allergic reactions, compared to placebo. Using Xolair for food allergies can help risk major side effects in cases of accidental exposure.
Academic papers
Improving AlphaFold2 performance with a global metagenomic and biological data supply chain [Munsamy et al., bioRxiv, March 2024]
Why it matters: lack of biological asset traceability and poor database diversity are bottlenecks to unlocking value from nature. This has led to controversies in biopiracy and models trained on a narrow sliver of the world’s biodiveristy. Basecamp’s biological data supply chain has increased sequence diversity, leading to better performing protein structure prediction and docking models whilst at the same time addressing the issue of equitable benefit sharing.Less than 0.000001% of life on earth is captured in public sequence databases available for deep learning in biology such as UniProt, NCBI or MGnify. These publicly available datasets are cultivated in narrow laboratory conditions, which lack geolocation and environmental metadata. The lack of geolocation data for the source of biological assets combined with improper Access and Benefit Sharing agreements driven by the UN Convention on Biological Diversity (2011 Nagoya Protocol), has led to controversies around biopiracy and impediments of assets that could have been valuable to human and planetary health.
The team at Basecamp Research show that their global metagenomics and biological data supply chain address both the issue of equitable benefit sharing of sequence information and the large sequence diversity gap with the aim to improve deep learning models.
It has been established that models based on multiple sequence alignment (MSA) like AlphaFold2 perform less well on orphan proteins for which MSAs cannot be generated. Basecamp demonstrates in this paper that the performance of AF2 can be improved when the MSAs are supplemented with diverse sequences from Basecamp’s knowledge graph. The RMSD compared to crystal structures is reduced up to 80% based on predictions for targets in CASP15 and CAMEO. Ligand docking prediction performance was also improved when using the MSA-augmented target structure.
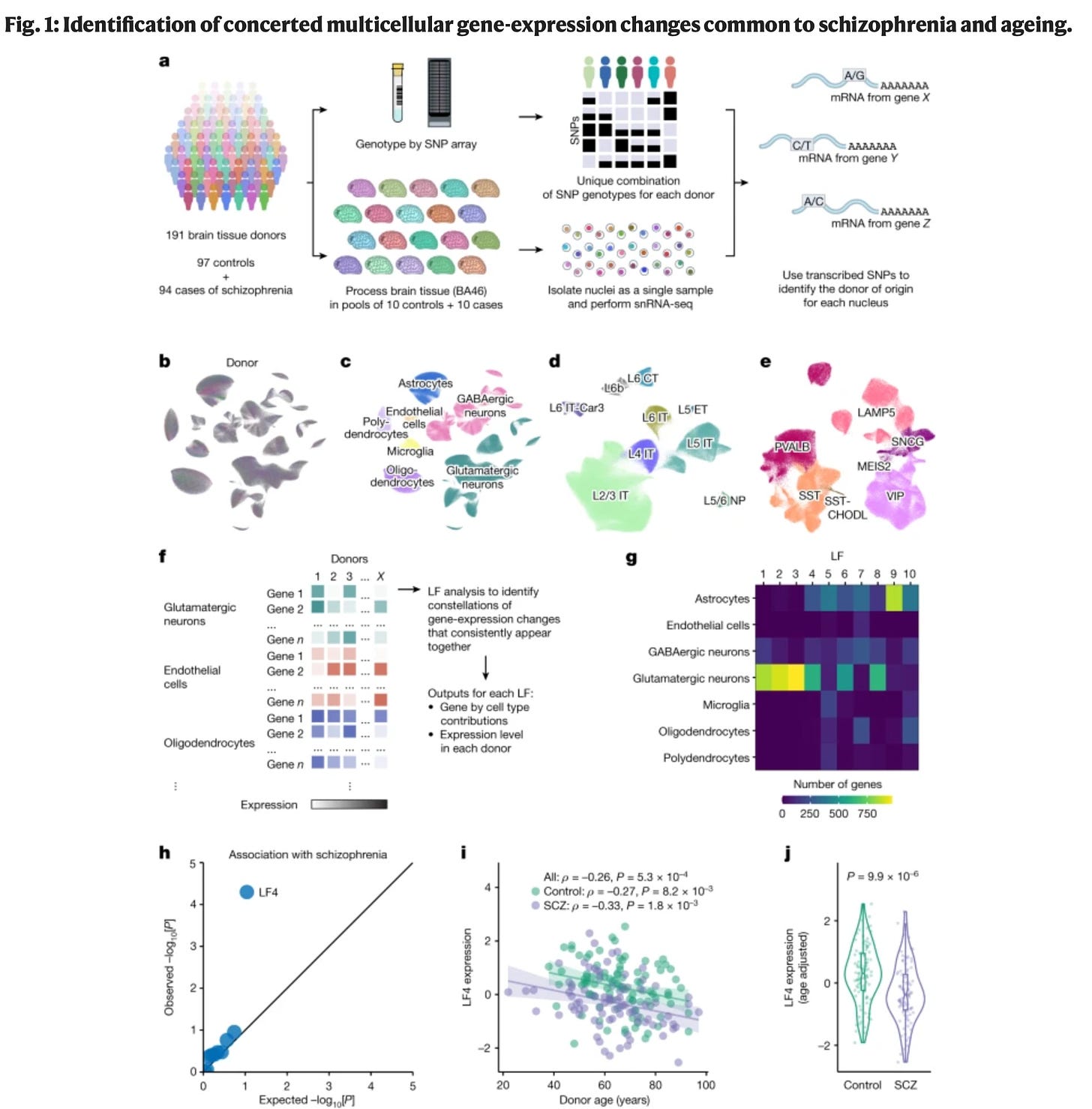
A concerted neuron–astrocyte program declines in aging and schizophrenia [Ling et al., Nature, 2024]
Why it matters: Researchers from the Broad Institute of MIT and Harvard have uncovered a coordinated network of gene expression patterns that is suggested to contribute to aging and schizophrenia, possibility offering a therapeutic mechanism to treat cognitive impairments of schizophrenia and help people maintain cognitive flexibility as they age.The researchers used single-nucleus RNA sequencing and machine learning to analyze gene expression data from 1.2 million cells from 94 people with schizophrenia and 97 older adults without schizophrenia.
This study is interesting as it focuses on the gene expression patterns of neurons as well as astrocytes at the same time, rather than just focusing on one cell at a time. By doing this, the researchers uncovered a highly coordinated gene expression program they have called SNAP: when neurons decreased the expression of certain genes related to synapses, astrocytes similarly changed expression of a distinct set of genes that support synapses. Scientists have long understood the role of neurons here, but the role of astrocytes had been less appreciated.
The team are working to now understand if these changes are present in other conditions such as bipolar disorder and depression.
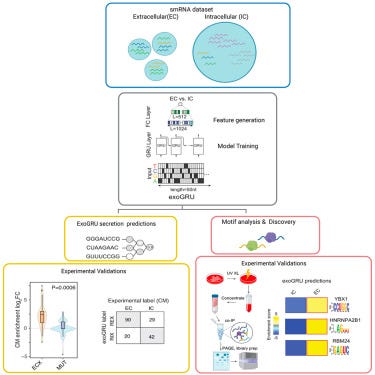
Revealing the grammar of small RNA secretion using interpretable machine learning [Zirak et al., Cell Genomics, 2024]
Why it matters: The Arc Institute team have developed a machine learning model that predicts RNA secretion. This research enhances our understanding of the complex processes involved, identifying the roles of various RNA-binding proteins, including both known and novel actors like YBX1, HNRNPA2B1, and RBM24, provides deeper insights into how RNA molecules are sorted and exported from cells, enriching our knowledge of cellular communication. The study's findings have significant implications for the development of new therapeutic strategies and the advancement of synthetic biology. By understanding the factors that govern RNA secretion, scientists can design synthetic RNAs with enhanced secretion capabilities, opening up possibilities for novel biomarkers and therapeutic interventions.The paper introduces ExoGRU, a machine learning tool designed to predict the secretion of small RNAs from primary sequences. This model accurately identifies small RNAs likely to be secreted, leveraging computational analysis to predict biological outcomes. The paper not only proposes a computational model but also validates its predictions experimentally. By analyzing the interactions between RNAs and RBPs in the extracellular space, the research confirms the model's accuracy and uncovers the grammatical rules governing RNA secretion.
A small-molecule TNIK inhibitor targets fibrosis in preclinical and clinical models [Ren et al., Nature Biotech, March 2024]
Why it matters: A recent paper and Phase I trial from Insilico Medicine demonstrates the power of an AI-first approach integrating multiple computational techniques - from AI-based target discovery using multiomics data to generative chemistry for drug design - to improve the speed and success rate of drug development. Using computational methods to identify a novel therapeutic target (TNIK) and to rapidly generate a potent and selective small molecule inhibitor of that target, the team was able to go from initial target discovery to demonstrating safety and efficacy of the drug candidate in phase I trials in ~18 months.Insilico Medicine just published one of the most impressive examples yet of end-to-end (i.e., target discovery through hit generation through lead optimization) AI-enabled drug development (figure below). The paper describes results from a Phase I trial in idiopathic pulmonary fibrosis (IPF), a disease of high clinical unmet need with only a couple approved drugs. PandaOmics, an AI-driven target discovery platform that integrates multiple computational approaches to analyze multi-omics data (human tissue samples, GWAS data, scientific literature, etc), discovered TRAF2- and NCK-interacting kinase (TNIK) as a top candidate for a novel target in IPF. While TNIK had been previously linked to cancer, Insilico discovered for the first time its connection to key fibrotic pathways and cell types.
Next, Chemistry42 (software for de novo small molecule design and optimization) was used to generate selective/potent drug candidates to TNIK. A lead candidate was identified, and in a series of in vitro and in vivo experiments, was found to exhibit impressive antifibrotic activity. In cell-based assays, it blocked TGF-β-induced fibroblast activation and myofibroblast differentiation, key drivers of fibrosis. In animal models, it attenuated fibrosis in the lung, kidney and skin, demonstrating potential to treat a range of fibrotic disorders. Notably, the compound also reduced inflammation, which often precedes and promotes fibrosis.
Finally, Insilico recently completed first-in-human phase I clinical trials in healthy volunteers. Two randomized, double-blind, placebo-controlled studies conducted in New Zealand and China evaluated the safety, tolerability and pharmacokinetics of the IPF drug. The results were encouraging, with the drug being generally well-tolerated with no major safety issues identified. The pharmacokinetic profile was favorable, with dose-dependent increases in exposure and a half-life suitable for once- or twice-daily dosing.
Going forward, Insilico’s IPF asset will be one of the most intriguing tests of the capabilities of AI-driven drug discovery. The drug is currently in a PhIIa trial. While IPF has been a challenging indication, recent positive PhIIa data from Pliant Therapeutics has galvanized the space.
Notable Deals
Partnerships
Merck and Pearl Bio partner to develop ‘smart’ biologics - Pearl Bio is a seed-stage start-up advised by George Church. The start-up has built a proprietary biomaterial platform to develop multi-functionalized biologics using non-standard amino acids. Although the upfront is not disclosed, the total deal value is $1B if the partnership progresses successfully. Pearl Bio claims their technology enables the era of ‘programmable’ biologics, with an ability to tune therapeutic properties such as half-life.
Merus and Gilead strike a deal to develop up to three novel trispecific antibodies that can harness the immune system to fight cancer. Merus will receive $56 million upfront plus a $25 million investment from Gilead.
Syngenta Crop Protection and Lavie Bio Forge Collaboration to Develop Biological Insecticidal Solutions. According to UN’s Food and Agriculture Organization, insects cause a $70B annual loss to global economy. The collaboration aims to harness Lavie Bio's unique technology platform for the swift identification and optimization of bio-insecticide candidates, combined with Syngenta’s extensive global research, development, and commercialization capabilities.
Financings
Boundless Bio seeks IPO - the San Diego biotech has developed a platform called Spyglass to seek out vulnerabilities in the cells carrying ecDNA and then designs small molecules to kill such cells. It has also developed an accompanying diagnostic. It has two assets in early stage testing with plans for data readouts in the next year. This IPO would be interesting as it would be only the seventh biotech in Phase 1 testing to price an IPO since the start of 2023.
Van Heron Labs raises $1.1 million seed to to enhance cell culture and utilization techniques using genomics, bioinformatics, artificial intelligence, and nanotechnology. The company’s mission is to develop the tools to power a sustainable bio-economy.
Relation Therapeutics has raised $35M from Nvidia’s NVentures, DCVC, Deerfield Management Company, Khosla Ventures, Abcam founder Jonathan Milner and ARK Invest. Relation is based in London and has built a platform based on genomics, machine learning in an integrated lab-in-the-loop between wet and dry to build the ‘largest’ single-cell bone atlas. The biotech plans to use this atlas to mine novel targets and figure out what type of drug to bring into clinical trials.
What we listened to
In case you missed it
A Blueprint for Breakthroughs [James Brodie]
What we liked on socials channels
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone