BioByte 070: NeuroAI, proteome-scale screening of protein degraders, open source biomedical LLMs, engineering better mRNA therapeutics
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. Happy decoding!
What we read
Blogs
The New Neuro AI [Nature Machine Intelligence, March 2024]
The relationship between neuroscience and AI is at a crossroads, with recent initiatives seeking to reignite the synergy between the two. Historically, neuroscience has inspired AI developments, from perceptrons to convolutional neural networks. Yet, as AI's focus has shifted towards complex architectures like transformers, the direct influence of neuroscience appears to have waned.
A panel at the Computational and Systems Neuroscience conference (COSYNE) reflected divergent views on NeuroAI's current influence and potential. Some experts remain optimistic about neuroscience's role in shaping future AI, suggesting that neuroscience could still unlock new AI methodologies. Others argue that while AI has significantly advanced neuroscience research, the reverse influence has been less impactful in recent decades.
Despite these debates, there's a concerted effort to foster NeuroAI through various platforms and academic programs, aiming to explore the shared principles of natural and artificial intelligence. Conferences like COSYNE and the Cognitive and Computational Neuroscience Conference (CCN) facilitate critical interdisciplinary dialogue, suggesting a fertile ground for collaboration.
The future of NeuroAI hinges on interdisciplinary engagement, with a growing need for scientists proficient in both neuroscience and AI. This emerging field promises to unravel the brain's computational secrets and pioneer smarter AI systems, provided it can harness the strengths of both domains.
Seeds of a wild idea [Tom Allen-Stevens, Direct Driller, March 2024]
Co-founders Ross Hendron and Professor Steve Kelly are pioneering the use of computational approaches to unlock the genetic potential of crops like wheat, soya, and maize by incorporating traits from wild plants that allow them to thrive in harsh conditions. One significant breakthrough is the editing of the wheat variety Cadenza to improve its photosynthetic efficiency by influencing the development of chloroplasts, aiming for a substantial increase in yield.
The venture, supported by £12 million in venture capital, operates from Milton Park, near Oxford, with a dedicated team working in advanced facilities.
Academic papers
Proteome-scale discovery of protein degradation and stabilization effectors [Poirson et al., Nature, March 2024]
Why it matters: Targeted protein degradation and stabilization are promising therapeutic modalities because they can target proteins that are considered “undruggable” and lack deep binding pockets. There are hundreds of E3 ligases in the human genome (a ubiquitin ligase that leads to degradation of the protein-of-interest), yet the majority of degraders in clinical development use only two (VHL and CRBN). By engineering a proteome-scale platform to functionally identify proteins that can promote the degradation or stabilization, a new study from Mikko Taipale’s lab at the University of Toronto identified many new degraders which were more potent and less sensitive to target identity and localization than commonly used E3 ligases like CRBN and VHL.Targeted protein degradation is a promising therapeutic modality that uses small molecules to induce interactions between a target protein and a degradation effector like an E3 ubiquitin ligase, leading to ubiquitination and proteasomal degradation of the target; however, current efforts have primarily focused on exploiting only a couple well-characterized E3 ligases such as cereblon and VHL.
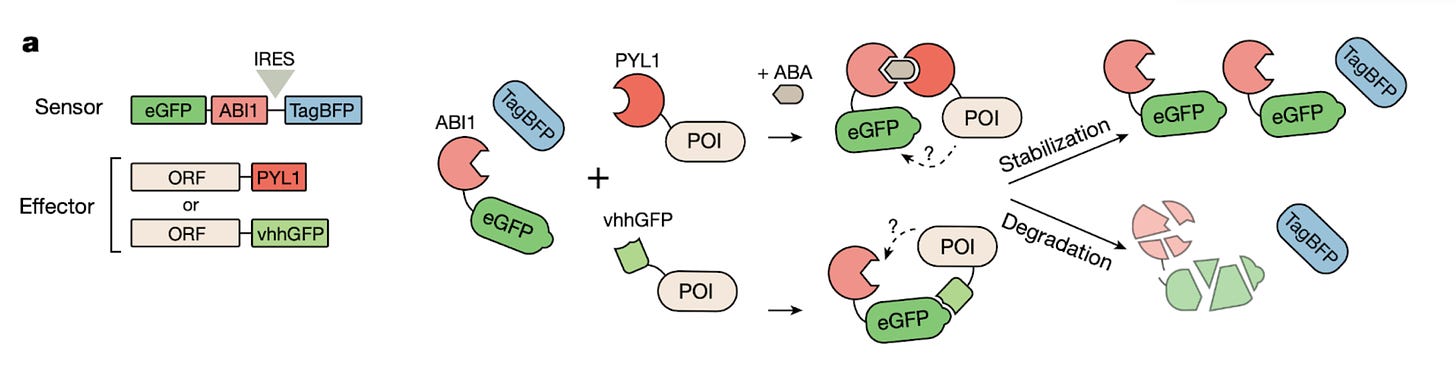
The researchers developed a screening platform to identify proteins across the human proteome that can degrade or stabilize a target protein in a proximity-dependent fashion (figure below). Large protein libraries were fused to proximity-inducing systems in cells expressing a GFP-tagged target protein, sorting the cells based on GFP levels to find hits. Hits were validated by testing their ability to degrade diverse target proteins, and evaluating the therapeutic potential of novel degraders in cell and animal models of chronic myeloid leukemia.
A large number of proteins beyond known E3 ligases were identified, including several that were more potent than the commonly used VHL and CRBN. Some effectors worked better on particular target proteins or cellular localizations (eg, ER-resident proteins were degraded most efficiently by membrane-associated effectors), however several top degraders were effective across a wide range of targets.
Branched chemically modified poly(A) tails enhance the translation capacity of mRNA [Chen et al., 2024]
Why it matters: Exogenous mRNA has proven effective in vaccinology; however, there has been limited success as a broad therapeutic modality given its intrinsic instability in the body and low translation efficiency. Here, the authors synthesized chemically modified mRNAs with multiple synthetic poly(A) tails, and demonstrate that multi-tailed mRNA had up to 19.5-fold higher luminescence post-transfection in vitro, and almost 2x greater signal detection in vivo. Using their constructs, the authors were able to achieve genome editing of Pcsk9 and Angptl3, demonstrating that capped branched mRNAs can be effective therapeutic modalities. mRNA therapeutics have largely floundered in clinic, due to instability (as they are susceptible to RNAses), and low translational efficiency; overcoming these often requires higher doses, which can lead to cytotoxicity. Although groups have tried several forms of mRNA modification - such as replacement of uridine with N1-methylpseudouridine to decrease immunogenicity, or circularizing RNAs to limit exonuclease degradation – these come with tradeoffs, such as reduced translation efficiency.
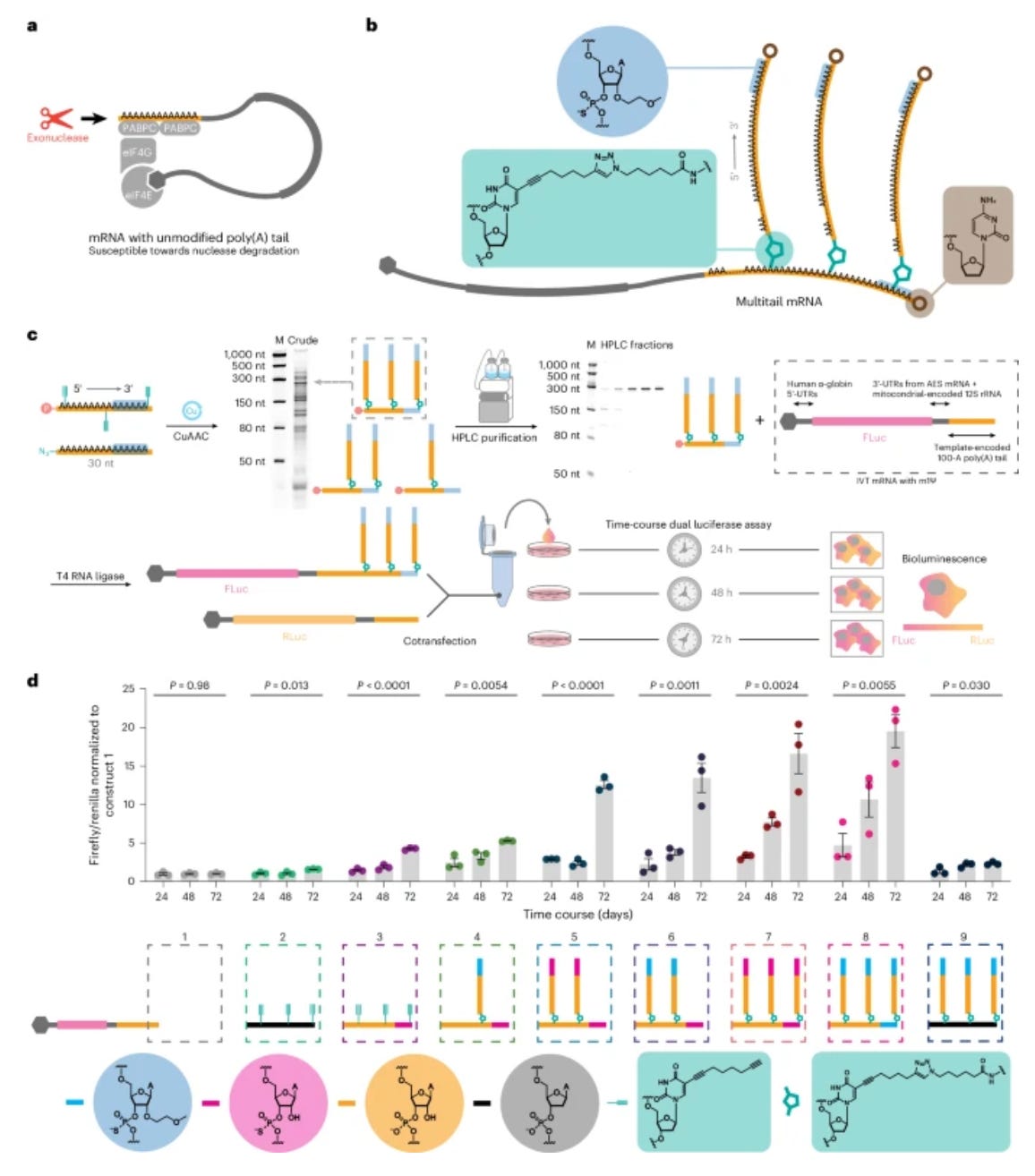
The rate-limiting step of translation appears to be the assembly of the initiation complex (dependent on the 5’cap) and stabilization through interaction with cytoplasmic poly(A)-binding proteins (PABPCs) and poly(A) tail. Here, PABC1 forms a multimeric protein complex when binding poly(A), generating a stabilized macrostructure. Additionally, the authors previously showed that introducing site-specific exonuclease-resistant modifications at the end of polyA tails increases mRNA stability and protein production; thus, they reasoned that multimerizing the poly(A) tail with branched structures and nuclease-resistant modifications would prevent RNA decay while maintaining the stabilized macrostructure to improve translation.
The authors generated several chemically modified, capped, and branched mRNA-oligo conjugates with multimeric poly(A) tails and evaluated how different chemistries and architectures affected mRNA translation. Indeed, they found that their multitail mRNAs led to extended protein expression in vitro and in vivo, without increasing immunogenicity. Of note, this allowed the delivery of proteins, such as Cas9, at significantly lower mRNA dosage (3 orders of magnitude), which holds promise for widening the therapeutic window of mRNA therapeutics. This work is exciting as it opens up an entirely new aspect of mRNA chemistry that has the potential to overcome the limitations of this modality to date.
BioMistral: A Collection of Open-Source Pretrained Large Language Models for Medical Domains [Labrak et al., arXiv, February 2024]
Why it matters: The development of BioMistral addresses critical challenges in integrating LLMs into biomedicine by offering an open-source, domain-specific model that rivals proprietary alternatives in performance.The multilingual capabilities of BioMistral pave the way for its application in diverse linguistic and cultural contexts, broadening the scope of its utility and impact.BioMistral introduces an open-source Large Language Model (LLM) tailored for the biomedical domain, leveraging the Mistral model and pre-trained on PubMed Central. It exhibits superior performance on ten established medical question-answering tasks compared to existing open-source medical models, with the added advantage of being lightweight through quantization and model merging techniques. This advancement marks the first large-scale multilingual evaluation of medical LLMs, offering datasets, benchmarks, and models for public use. BioMistral's construction involves pre-training on a carefully curated dataset, employing advanced model adaptation, merging strategies, and quantization techniques to optimize performance across multilingual contexts.
BioMistral underwent comprehensive evaluation across a benchmark of ten medical question-answering tasks in English and seven other languages. The evaluation showcased BioMistral's exceptional performance, particularly in English, and highlighted its competitive edge against both proprietary and open-source counterparts. Model merging techniques like SLERP, DARE, and TIES further enhanced capabilities. The study also explored the model's multilingual potential, demonstrating robust performance across different linguistic contexts despite the challenges of translation quality. Quantization techniques enabled the model to be more accessible on consumer-grade devices without significantly compromising its performance.
Generative AI for designing and validating easily synthesizable and structurally novel antibiotics [Swanson et al., Nature Machine Intelligence, March 2024]
Why it matters: In 2019, ~5m deaths were associated with drug-resistant infections and projected to grow to 10m by 2050 as antimicrobial resistance continues to outpace the discovery of new antibiotics. Swanson et al. demonstrate how a generative model can design new antibiotics that are bioactive against one the most virulent pathogens A. baumannii.Existing molecular prediction models can evaluate chemical datasets for antibiotic properties in order to discover new potential drugs. However, these models must evaluate each chemical one-by-one from existing chemical libraries, which prevents them from exploring vast chemical spaces in reasonable time and are unable to generate truly new chemical matter.
To overcome these challenges, the authors built a generative model, named SyntheMol, to generate novel chemistry with the desired properties without evaluating chemicals one-by-one. In order to increase synthesizability rates, SyntheMol uses 132,000 molecular building blocks with known reactivities and 13 well-validated chemical synthesis reactions. Which translates to 30 billion molecules that are easy to synthesize at 80% success rate.
In this paper, SyntheMol was trained to design molecules against A. baumanii. They synthesized and validated 58 of the generated molecules; with 6 structurally diverse molecules displaying potent antibacterial activity.
A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche [Zepeda-Rivera et al., Nature, March 2024]
Why it matters: This study reveals that a bacteria that normally lives in the mouth was found in 50% of colon cancers, which supports prior research on the interactions between the gut microbiome and colon cancer. As rates of colon cancer increase, particularly in younger people, the study sheds light on how bacteria can be used in colon cancer treatments. The researchers used a combination of classical microbiology culturing, whole genome sequencing, and comparative genomics to analyze Fusobacterium strains from colorectal tumors as well as the oral cavity. They found that a specific clade of Fusobacterium nucleatum (Fna C2) is found to dominate the colorectal cancer tumor niche while Fna C1 is found in the oral cavity. They then used various functional studies to find that Fna C2 is associated with increased metabolic potential, colonization of the gastrointestinal tract, and intestinal adenomas in mice. Fna C2 is also found to be more prevalent in the stool of patients with colorectal cancer compared to those without.
These findings suggest that Fna C2 plays a significant role in the development and progression of colorectal cancer and has implications for diagnosis and treatment. Now that the presence of bacterial populations in colorectal cancer is confirmed, the next challenge is understanding whether and how the microbes contribute to disease.
Notable Deals
M&A:
Novo Nordisk to acquire RNA biotech Cardior for 1B EUR - the deal includes Cardior’s Phase II therapy in phase 2 development for heart failure. The move comes as the Danish pharma seeks to strengthen its cardio pipeline
Financings:
Nkarta targets $244M raise and switches focus from CAR-NK in cancer to autoimmune cell therapy
Taxa Technologies raises $2.5M pre-seed to leverage microbiome technology to create better personal care products
Strategic:
What We Listened To
In case you missed it
Ming Tommy Tang: On A Mission To Teach 1M People Bioinformatics
What we liked on socials channels
Events
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone