BioByte 074: Why Mars is tough for life, an autoantibody signature for MS, selective compartmentalization for proteins, AI-designed CRISPR, RNAi-based drug design
Welcome to Decoding Bio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. Happy decoding!
Welcome to Spring. We know that Spring Cleaning is tough, but at Decoding we have you covered with our tried and true routine. Step 1: Remove all paintings and artwork around your house. Step 2: Replace them with pages from Decoding. Step 3: Sit in silence as you realize, once again, that knowledge transfer through osmosis is real and that you’ve been learning the wrong way your entire life.
What we read this week
Why Nothing Can Grow on Mars* (*Probably) [Devon Stork, Asimov Press, 2024]
Populating Mars has pervaded popular culture for more than a decade, with Elon Musk stating a personal goal of enabling human settlement of Mars and movies such as The Martian hitting the cinemas. But clear five clear challenges remain: (1) Water: Life as we know it requires liquid water for survival, Mars has little available water - most of it is frozen (-100°C) in its poles or has extreme salt contents. (2) Radiation: Mars receives 1000x more UV than Earth (3) Toxins: soil consists of high levels of perchlorate, a chemical toxic to humans (4) Temperatures: whilst temperatures at the equator can reach 20°C, the poles can plunge to -150°C. (5) Atmosphere: The Mars atmosphere is composed of 95% CO2 and nearly devoid of oxygen.
However, synthetic biology may offer an option to build sustainable life in these extreme conditions. There are already examples of microbes that resist each of these challenges individually. The difficulty is that there is no known organism that can thrive with all 5 challenges at once. But by combining ‘extremophilic traits’ found in organisms across Earth, biologists may be able to create a hybrid microbe that can thrive in the harsh environment of Mars.
An autoantibody signature predictive for multiple sclerosis [Zamecnik et al., Nature Medicine, 2024]
Why it matters: Multiple sclerosis (MS) is a debilitating, chronic autoimmune disease that is canonically marked by demyelination of neurons throughout the brain and spinal cord. Although traditionally believed to be a T-cell mediated disease, the efficacy of B-cell depleting therapies has revealed the critical role of these cells and antibodies in the progression of MS. In recent years, it has been hypothesized that autoantibodies may play a role in MS, but this has been difficult to elucidate. Here, the authors retrospectively profiled a cohort of 10 million individuals to generate whole-proteome autoantibody profiles of MS patients. They identified a common pattern of autoantibody reactivity to an epitope similar to that found in Epstein-Barr Virus that is present years before developing MS symptoms, and they have higher levels of serum neurofilament light (sNfL - a marker of axonal damage). This pattern is present for years, and represents a potential signature for clinical use in the diagnosis of MS.The authors profiled a prospective incident MS cohort of 250 patients from the Gulf War era (GWEMSC) that encompassed over 10 million individuals in the active duty US military population. Serum was pulled from multiple periods: 1) earliest available from entry into active duty, 2) ~5 years prior to their first clinical manifestations, and 3) another sample roughly 1 year after the first attack. They used a whole-human proteome seroreactivity technique called Phage Immunoprecipitation Sequencing (PhIP-Seq) where proteome-scale peptide libraries are displayed on bacteriophages and subsequently immunoprecipitated using an individual’s antibodies before being analyzed via high-throughput DNA sequencing. They found that 10% of MS patients had developed number of autoantibodies years before their diagnosis; these autoantibodies largely bound to epitopes similar to those found in many infectious agents - including two EBV proteins (the association between EBV and MS has become increasingly well known in recent years). Additionally, these patients had elevated levels of neurofilament light (a marker of neuronal damage).
This work reveals the presence of prospective autoantigen-specific biomarkers years before the manifestation of MS symptoms, and confirms that neuronal injury is present in this preclinical phase. Although significant work is necessary to decipher additional dynamics of pathophysiology, this study is a critical addition to understanding the development of MS and how to identify subsets of patients prior to the onset of symptoms.
Chemical codes promote selective compartmentalization of proteins [Kilgore et al., bioRxiv, April 2024]
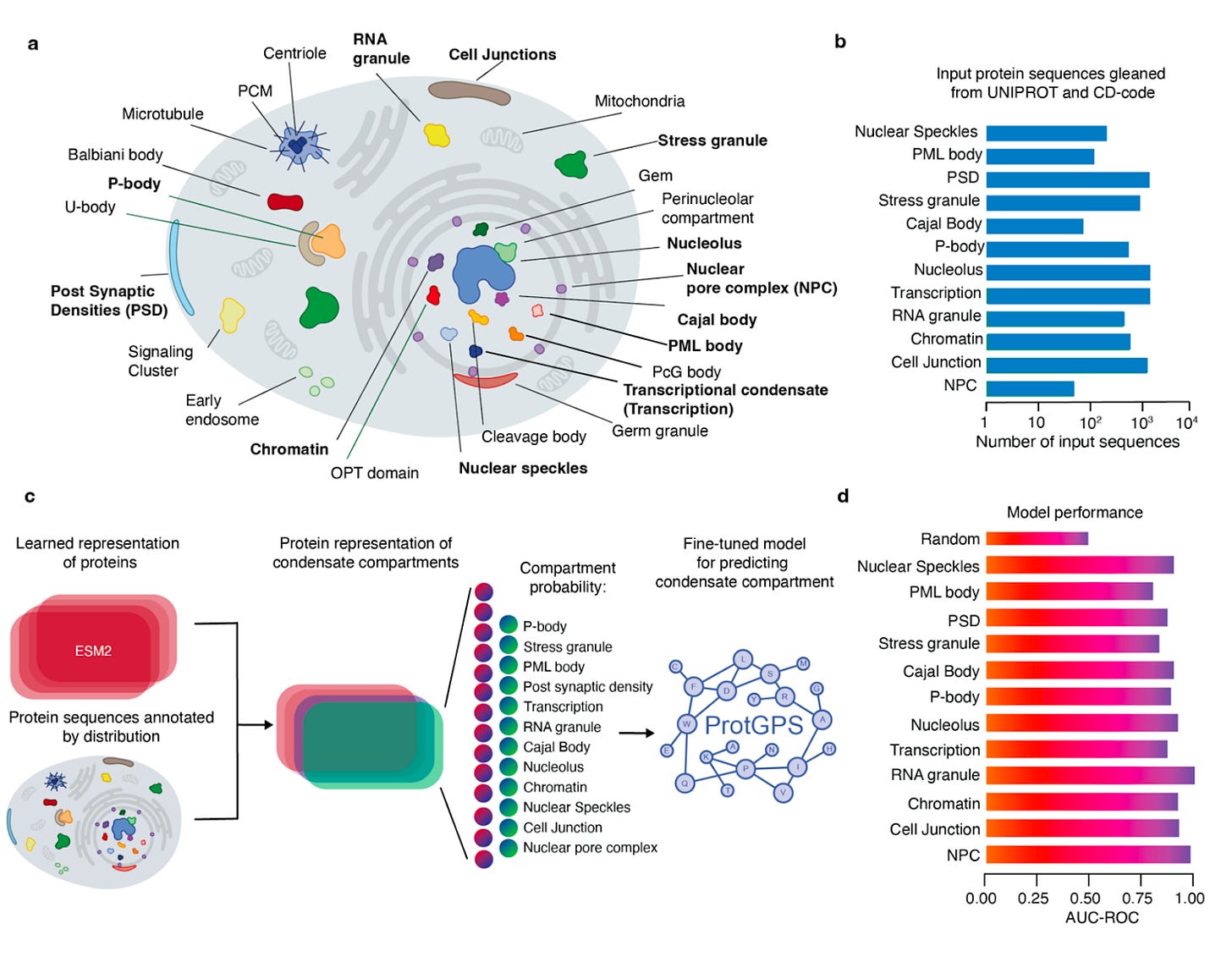
Why it matters: In this paper, the authors demonstrate that protein sequences not only contain the folding code, but also the chemical code governing their distribution in specific cellular compartments. If this code is solved, it could have important implications for our understanding of cellular function and dysfunction.Kilgore at al. developed ProtGPS, a LLM that was adapted from ESM2, which can predict the compartment localization of human proteins and generate novel protein sequences that selectively assemble into specific compartments in cells.
In order to fulfill their physiological function, proteins with shared functions must correctly assemble at specific locations in the cell. Shape complementarity among structurally stable portions of proteins have dominated the model of protein assembly, but more recently, evidence has pointed to the fact that large assemblies of proteins (aka condensates) also occur through weak multivalent electrostatic interactions.
In another study we covered, it was demonstrated that condensates can harbor distinct chemical environments, suggesting they have different solvent proteins and thus leads to the hypothesis that proteins with shared physiological functions assemble due to a shared chemical salvation code.
Using more than 5,000 protein sequences annotated for 12 condensate compartments to train the model, ProtGPS computes the probability of a protein being found at a given compartment. The AUC-ROC curve showed an accuracy of 0.83-0.95.
The authors also generated novel proteins designed to assemble in particular compartments and then tested them in human cells, which showed remarkable accuracy. Using ProtGPS, the team could also determine if pathogenic mutations would lead to changes in subcellular distribution.
Design of highly functional genome editors by modeling the universe of CRISPR-Cas sequences [Ruffolo et al., bioXriv, APril 2024]
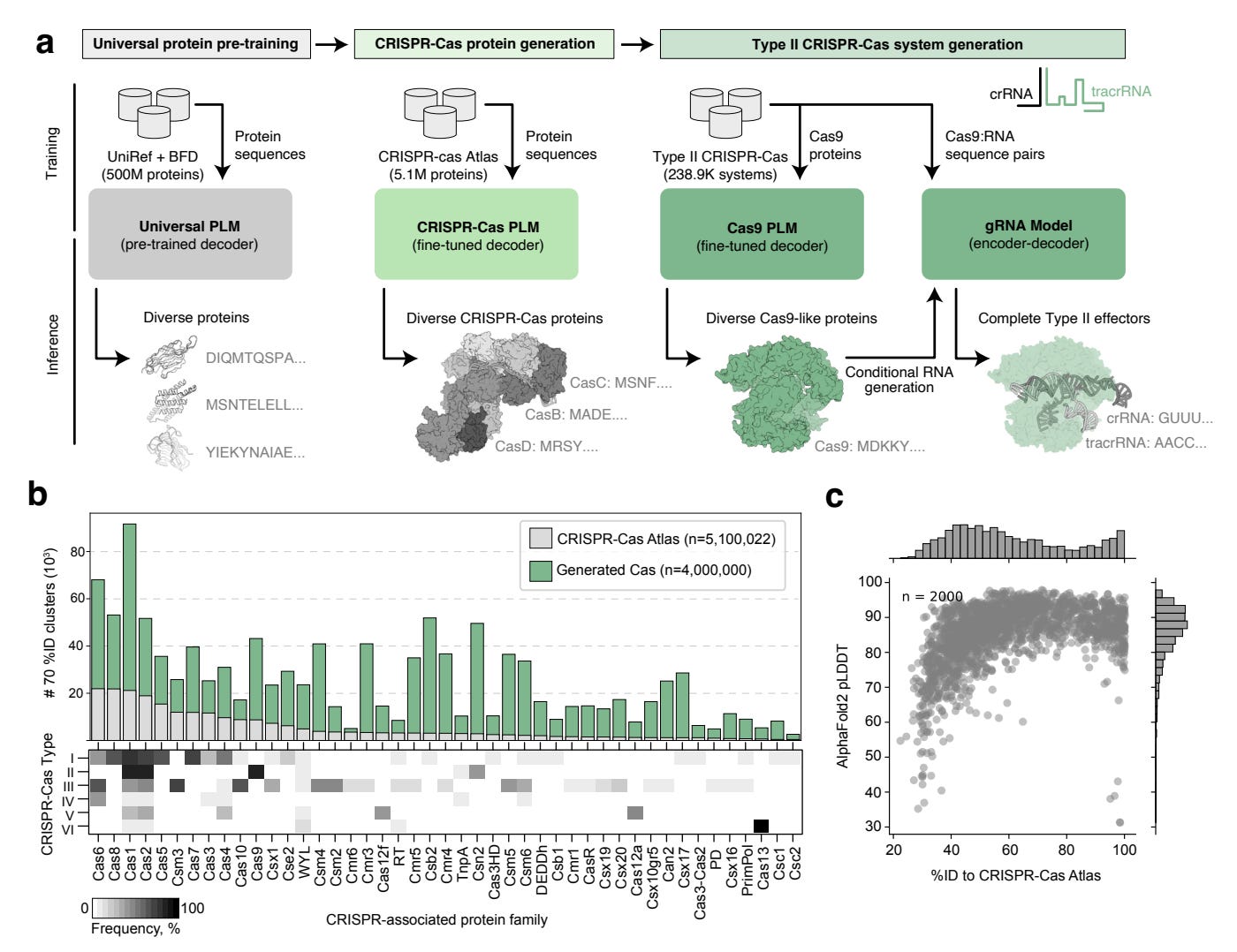
Why it matters: The development of AI-designed CRISPR systems such as OpenCRISPR-1 is a game-changing advancement for multiple reasons. Firstly, it represents a leap in precision medicine, offering the potential for highly targeted therapies with minimized off-target effects, which is crucial for the safety and efficacy of gene therapies. Secondly, the ability to bypass evolutionary constraints and design novel proteins with desired functionalities could accelerate the development of treatments for diseases that are currently difficult or impossible to treat. Additionally, the open release of OpenCRISPR-1 encourages global collaboration and innovation, broadening the potential applications of this technology in medicine, agriculture, and biotechnology. This progress underscores the merging of computational power and biological science, promising to usher in new frontiers in genetic research and therapy.A recent study by the team at Profluent highlights a significant breakthrough in CRISPR-Cas genome editing technology, achieved by integrating artificial intelligence (AI). Researchers have successfully developed a new gene editor, OpenCRISPR-1, using large language models (LLMs) that were trained on a comprehensive dataset of over one million CRISPR operons. This novel editor shows improved activity and specificity compared to the widely-used SpCas9, despite being significantly altered in its protein sequence.
This advancement is particularly notable as it demonstrates the use of AI to transcend natural evolutionary limits, creating gene editors with optimized features that were not previously achievable. The AI-designed OpenCRISPR-1 not only functions effectively in human cells but also exhibits compatibility with base editing technologies, expanding its potential utility in therapeutic settings.
RNAi-based drug design: considerations and future directions [Tang et al., Nature Reviews Drug Discovery, April 2024]
Why it matters: A recent Nature Rev Drug Discovery review highlights the progress and potential of RNAi therapeutics in treating diseases caused by abnormal gene activity. Key priorities for the field include siRNA design to improve stability and immunogenicity, delivery to specific tissues and cell types of interest, and expanding siRNA applications to new areas such as isoform-specific modulation, antiviral therapies, long non-coding RNA targeting, and combination therapies.A recent review published in Nature Reviews Drug Discovery surveys the field of RNA interference (RNAi). RNAi is a process in which small pieces of RNA, called small interfering RNAs (siRNAs), can silence specific genes by degrading their mRNA before it can be translated into proteins.
Scientists have learned to design synthetic siRNAs that can target virtually any gene. Turning siRNAs into effective drugs, however, has been challenging (see the history of the modality in the timeline below). Early siRNAs were quickly degraded by enzymes in the body and triggered immune responses. To overcome these issues, researchers have developed chemically modified siRNAs that are more stable and less immunogenic.
Another key challenge has been delivering siRNAs to the right tissues. Lipid nanoparticles (tiny fat droplets) and GalNAc conjugates (siRNAs attached to a sugar molecule that binds to receptors on liver cells) have enabled successful delivery to the liver, leading to FDA-approved treatments for liver-related genetic diseases. Scientists are now exploring new delivery strategies using antibodies, peptides, aptamers (RNA or DNA molecules that bind specific targets), and lipophilic molecules to reach other tissues.
A few exciting future directions are emerging:
The development of siRNAs that can selectively silence specific gene isoforms. Isoforms are different versions of a gene that arise through alternative splicing. In some cases, certain isoforms may be more disease-related than others, and by targeting specific isoforms, there is a lower risk of off-target effects.
RNAi therapeutics for viral infections. siRNAs can be designed to target essential viral genes, potentially blocking viral replication and spread. By targeting conserved regions of the viral genome that are less likely to mutate, scientists could develop siRNA drugs that remain effective against a broad range of viral strains, including newly emerging variants. This approach could be particularly valuable for combating viruses that rapidly evolve to evade conventional treatments, such as influenza and HIV.
Long non-coding RNAs (lncRNAs) represent another exciting frontier for RNAi therapeutics. These enigmatic RNA molecules, once dismissed as "junk" DNA, are now known to play crucial roles in regulating gene expression and various cellular processes. However, the complex and often cell-type-specific functions of lncRNAs pose unique challenges for siRNA design and delivery.
Notable Deals
Partnerships
Ochre Bio and Boehringer Ingelheim form a $1.3B biobuck research pact for chronic liver disease - the partnership includes $35M upfront to develop treatments for chronic liver disease that aim to harness the organ’s capacity to regenerate.
Deciphex and Novartis collaborate for AI in preclinical studies - The collaboration seeks to develop a suite of AI tools that streamline the pathology evaluation of toxicological and efficacy studies.
Financings:
Xaira Therapeutics’ $1B bet to transform drug discovery using generative AI - the Seattle-based biotech has a star studded team with researchers from David Baker’s Institute of Protein Design and CEO leadership from Marc Tessier-Lavigne (former CSO Genentech).
Regeneron commits $500M to a new fund - Regeneron Ventures aims to build and grow biotech companies and Regeneron will be the fund’s exclusive limited partner. The $500M will be deployed over the next 5 years.
Corner Therapeutics launches with $54M to create multi-purpose immunotherapies for cancers, influenza and HIV.
What we listened to
What we liked on socials channels
Field Trip
Did we miss anything? Would you like to contribute to Decoding Bio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @ketanyerneni @morgancheatham @pablolubroth @patricksmalone