Welcome to Decoding TechBio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. Subscribe for free to read our weekly review and deep dives on areas we’re excited about. For more on us, check out our mission statement here. If you’d like to connect or collaborate, please shoot us a note at decodingtechbio@gmail.com. Happy decoding!
Happy Monday! Welcome to Fall —get cozy and buckle up. Whether PSL stands for Photostimulated Luminescence or Pumpkin Spice Latte, we’ve got you covered.
We were delighted to hear from so many old friends and new readers after publishing our first weekly piece last week. As a writing collective, we strive to learn and build in public and in partnership with our readers and supporters across academia and industry. Thanks for your support. Now, for what we consumed this week!
What we read
Academic Papers
In silico QSAR modeling to predict the safe use of antibiotics during pregnancy [Drug and Chemical Toxicology]
One of the greatest opportunities for in silico methods is to study populations that are challenging to research in vivo. Pregnant populations are a great example, where much of our current understanding of teratogenicity, or the ability of a compound to cause defects in a developing fetus, stems from anecdotal evidence and retrospective studies.
Researchers recently applied in silico approaches to develop a model that predicts safe use of antibiotics (one of the most commonly prescribed drugs) in pregnancy using Quantitative Structure-Activity Relationship (QSAR), a well-known in silico approach that correlates chemical structures of molecules with the toxicity of endpoints (great two-part lecture here). In lieu of animal or other human models, this paper demonstrates the feasibility of developing robust and high-performing in silico models for understanding toxicity in pregnancy.
A tissue-engineered artificial human thymus from human iPSCs to study T cell immunity [Nature]
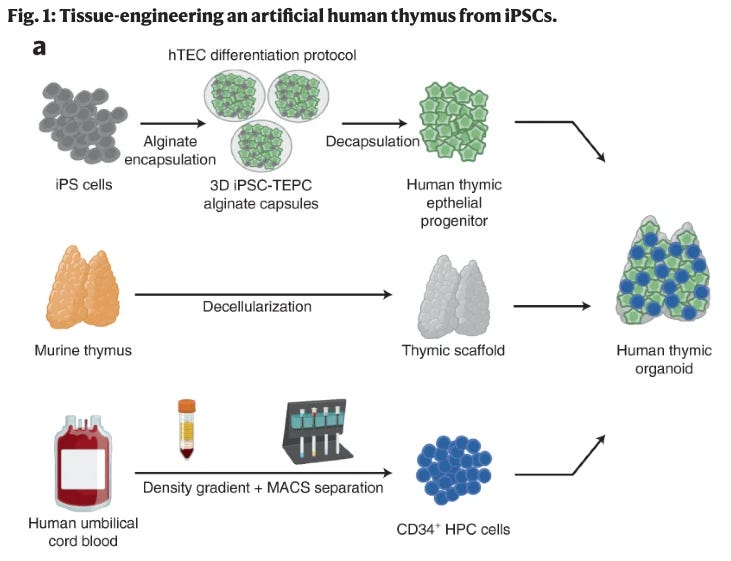
We’ve engineered a thymus! Fascinating bioengineering work in this recent piece leverages a tissue engineering method using a 3D scaffolding and enables the generation of an artificial human thymus from inducible pluripotent stem cells (iPSCs). What’s a thymus and why do we care about it? This picture appreciates the thymus in all its glory.
The thymus is a primary lymphoid organ of the immune system where T cells mature. Being able to develop a human thymus in vitro unlocks a whole host of research opportunities for studying T cell maturation and specifically, adaptive immune responses. We’re excited about the next iterations of this work that could explore developing an artificial human thymus using a patient’s own iPSCs. Here’s how it all happens:
Blogs
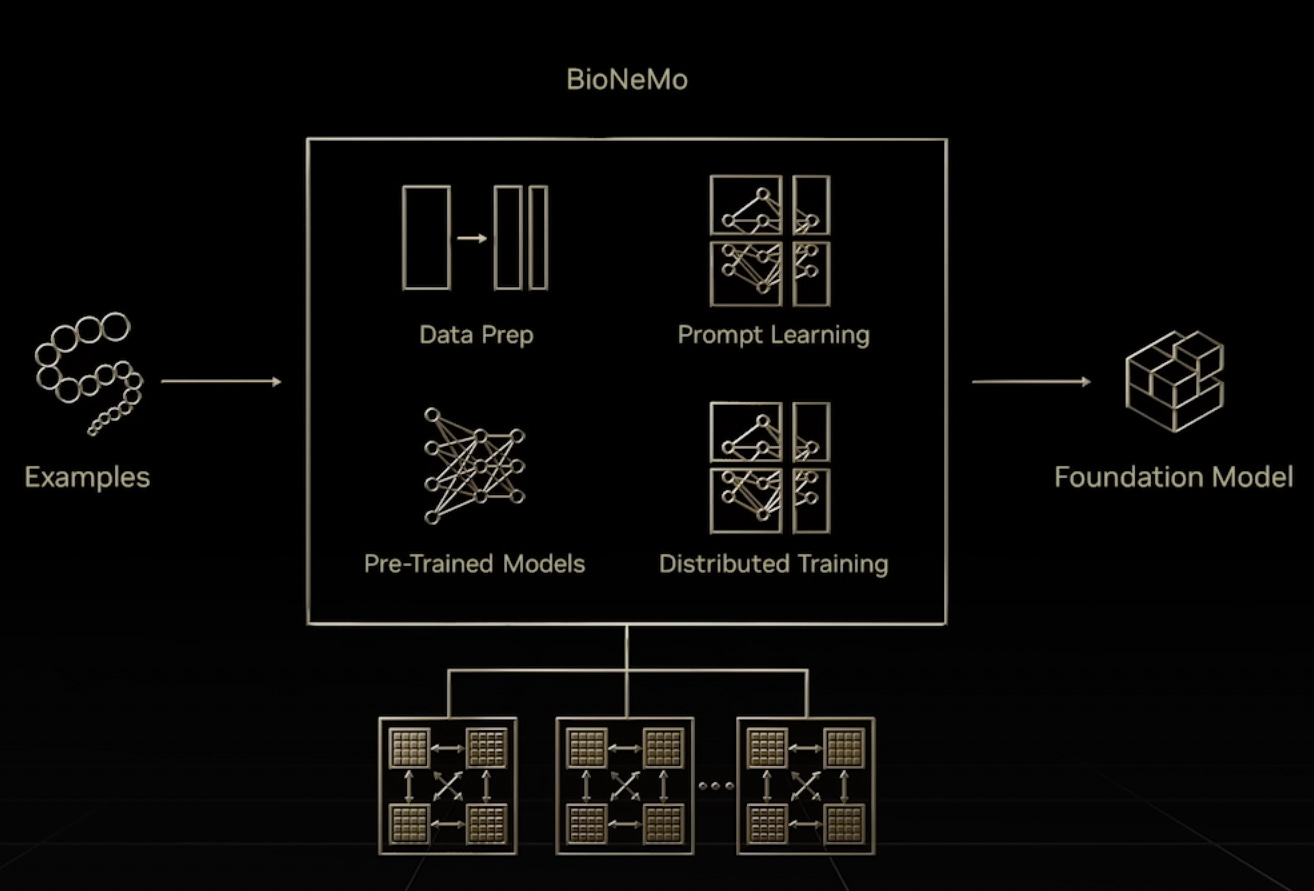
NVIDIA Expands Large Language Models to Biology
Advances in compute are being increasingly translated across the tech world to the biotech world. NVIDIA’s BioNeMo is a large language model (LLM), based on transformer architecture, that can learn in an unsupervised fashion without labeled data-sets. BioNeMo will support the language of life – biology and chemistry – and enable the next wave of data-driven therapeutic discovery.
Machine learning powers biobank-driven drug discovery [Nature News]
Over the last two decades, human genetic data has played an integral role in supporting emerging drug programs by supporting population-scale surveys known as genome-wide association studies (GWAS), which compare genetic features of patient cohorts to those of healthy controls. Harnessing genomic data from research biobanks, researchers at startups and pharmaceutical companies have demonstrated an insatiable appetite for clinicogenomic data, and some of these approaches are starting to bear fruit.
In April, BioAge Labs, a growth stage TechBio company leveraging machine learning on biobank assets from prior aging studies, dosed their first patient for a muscle atrophy program. Pharma companies such as Amgen have paid premium prices for access to biobanks over the last several years and groups like Regeneron are partnering with leading clinicogenomics research institutions to instantiate large-scale genetic sequencing projects.
As more focus and investment turn to biobanks, we’ll be following the productivity of these approaches and how companies navigate nuances of this challenging terrain from validating sample quality to managing patient consent.
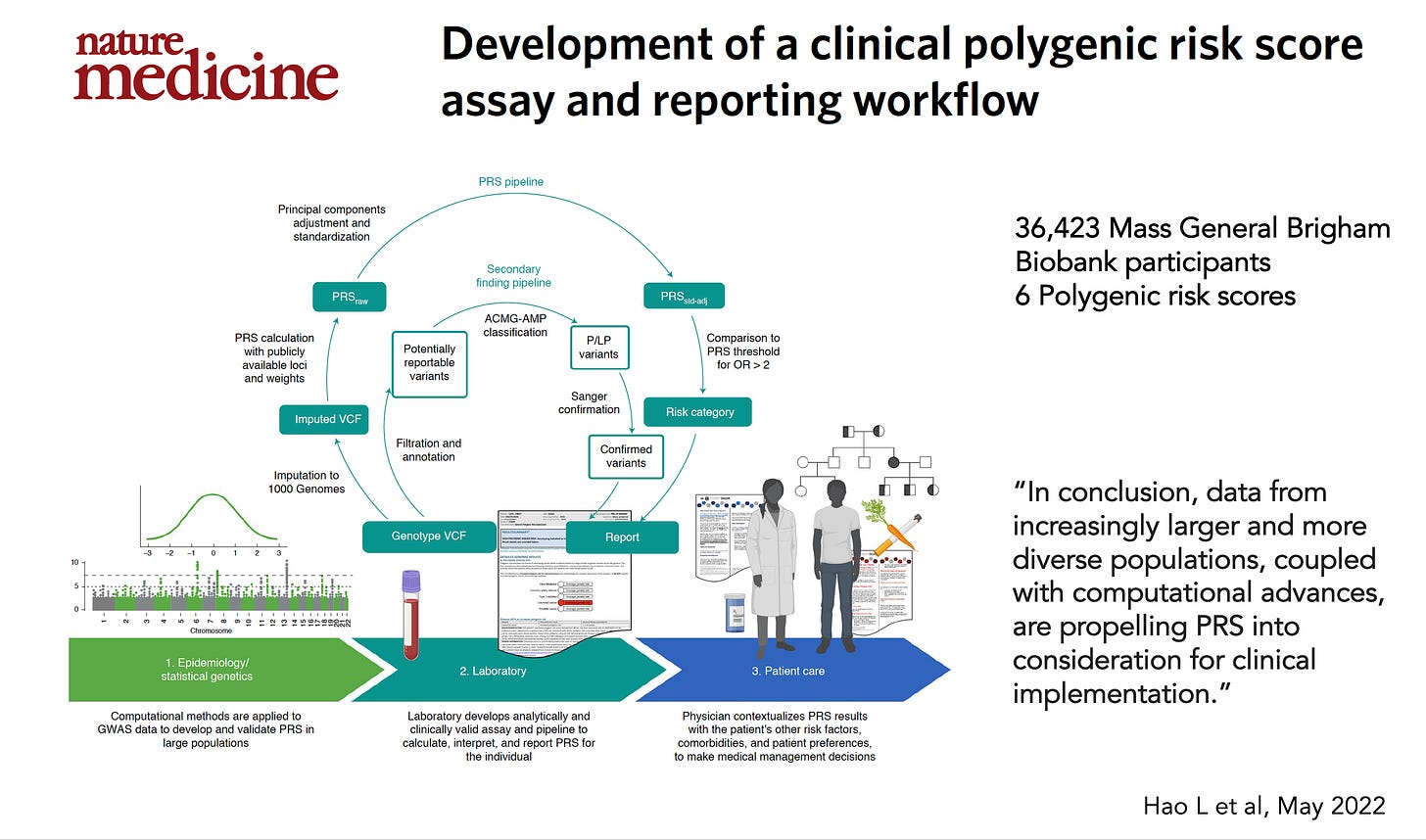
Human Genomics vs. Clinical Genomics [Dr. Eric Topol]
We loved this brief piece penned by Dr. Eric Topol, which speaks to the great need to drive adoption of genomics insights in the form of polygenic risk scoring in clinical practice. In addition to the first post, he published a thoughtful addendum that clarifies where the gaps are in routine medical practice.
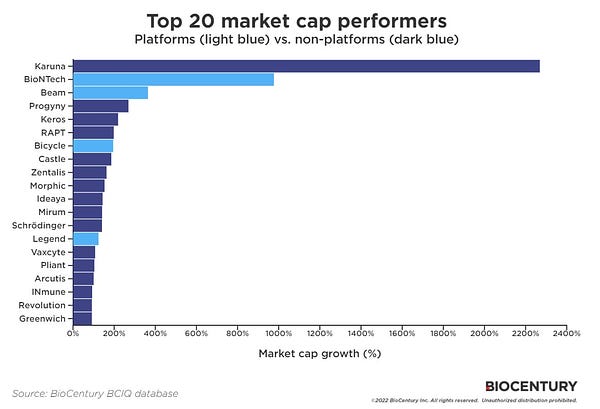
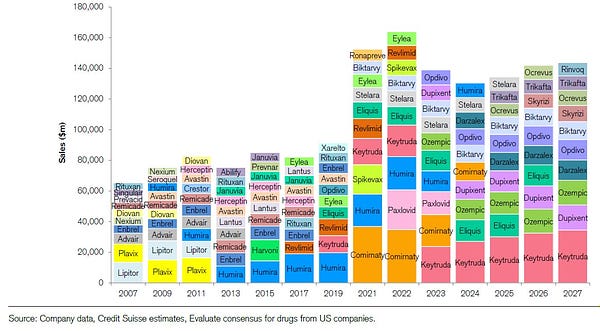
Deep-dive into platform biotech companies: phenotypes, business models, destinations... [Biodraft]
Liang Chang and Kirill Karlin dissect the anatomy of platform biotechs. They dig into what are platform biotech companies and their business models? What are some of the critical strategic decisions in these companies’ lifecycles? What are the outcomes if things go well, or not well? And many more questions.
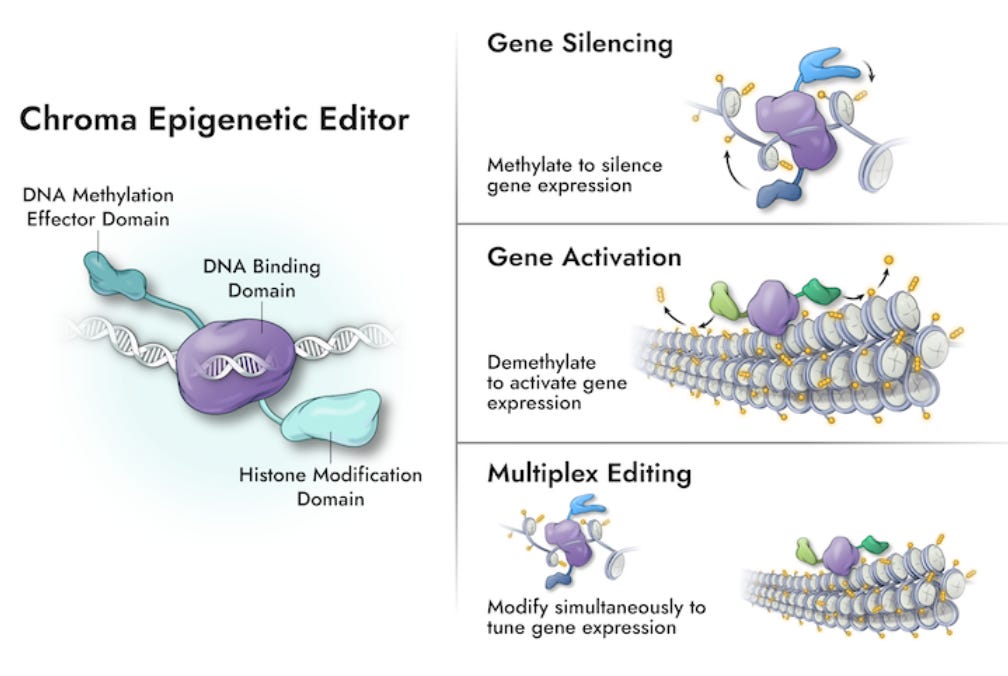
Chroma Medicine and Tune Therapeutics: Two companies take up epigenome editing [Nature News Feature, Sept 2022]
Epigenome modification as a therapeutic mode of action has been utilised since the 90s through small molecules that act as therapeutic hammers, rather than scalpels. The two companies described in this article, Chroma and Tune are developing precise epigenome editors to manipulate DNA expression without irreversibly mutating it. Both companies use ‘dead’Cas or dCas bound to different transcriptional activators, repressors or ‘writers’. Whilst it is still early days for these companies, the CEOs and scientific founders behind the technology say that “there have been no ‘Oh no’ moments yet.”
Opinion: The need for a genomics-driven pathogen global warning system [Phil Febbo, Devex, 2022]
In the past 5 months, we’ve seen two new outbreaks: monkeypox and polio. The technology for genomic surveillance is here; to prevent this, we require better surveillance networks. Dr. Phil Febbo, Chief Medical Office at Illumina, proposes what he names the ‘Global Pathogen Surveillance Network’: a collaboration between the US CDC, WHO, health systems, the defense sector, and the private sector to bring critical capabilities and resources to the table to prevent and control new pathogen outbreaks.
What we listened to


What we liked on Twitter
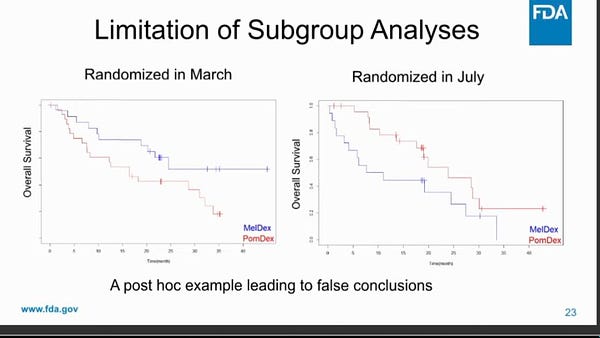
A friendly reminder that there are significant limitations of subgroup analysis! False positives, false negatives, and limited ability to inform and guide individual treatment decisions:



Where we went (and are going)
DeSci Boston
Well, we missed the DeSci Boston, a 1-day event to bring together web3 developers, DeSci leaders, and funders in the DeSci ecosystem, last week, but we heard it was a good time. Did you attend? Share your thoughts with us, we’d love to highlight what our community learned. Follow the DeSci Community Eventbrite page for more information about upcoming events.
2022 Founder-led Biotech Summit
We signed up for the 2022 Founder-led Biotech Summit hosted by our friends at Pillar VC. Learn more about this upcoming virtual event here.
Private financings this week
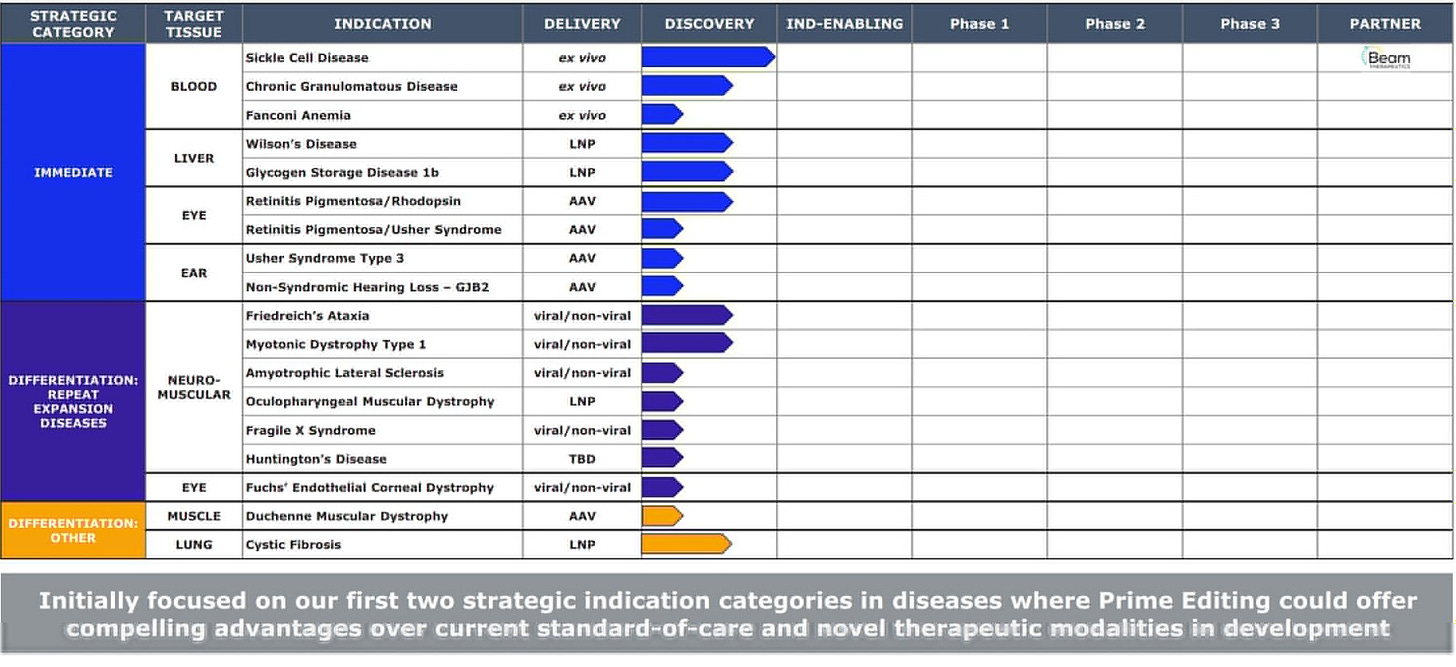
Prime Medicine filed for an estimated $200M IPO, just over 2 years after it started operations. Interestingly enough, they’re going public with their entire pipeline (18) in the discovery stage. Not a single in IND-enabling. Rushed or calculated? Only time will tell.
Field trip
Did we miss anything? Would you like to contribute to Decoding TechBio by writing a guest post? Drop us a note at decodingtechbio@gmail.com or chat with us on Twitter: @ameekapadia @pablolubroth @patricksmalone @morgancheatham @ketanyerneni