BioByte 012
Churchisms, generative diffusion models for proteins, clinical trials are real expensive (confirmed!), lecanemab data, drag-and-drop genome insertion, smart bandaids, and more
Welcome to Decoding TechBio, a writing collective focused on the latest scientific advancements, news, and people building at the intersection of tech x bio. If you’d like to connect or collaborate, please shoot us a note here. Happy decoding!
Happy Tuesday! Hope the start to the holiday season has been treating everyone well, and hope that people haven’t been crying (too much) because of the World Cup. We’re excited to meet a ton of you during JPM - stay tuned for events galore as events and meetings get scheduled. Feel free to reach out to any of us directly if you’d like to meet!
What we read
Blogs
George Church [Dylan Neel and Henry Klingenstein, Biomarker, 2022]
Biomarker is a fantastic newsletter by Dylan Neel and Henry Klingestein that features interviews with prominent scientists, founders, and investors in biotech and medicine. Last week featured an interview with George Church. Some highlights:
Every grad student should know the work of Thomas Hunt Morgan who discovered the chromosome theory, but few do
Much of George’s work in sequencing was inspired by his passion for photography
On George’s diverse scientific interests: When approaching a new area, he is focused on doing one specific thing to find a wedge in, and does not try to learn the entire field. He learns by doing, and quickly finds a grad student in his lab to pursue the new topic. They purposely work on technology, rather than biology, so as not to disturb or threaten the scientists already working in the field. This way, the incumbents know that they will get a new technology to add to their toolkit, and then George will shift focus to another field.
On how to select a good research project: focus on enabling technologies that can be broadly applied across applications and industries. Aim for pushing a technological capability forward by an order of magnitude, not incremental progress that others can copy. Pick something that is beneficial for society, and not just cool to do.
An area of science outside of his direct field that George is excited about: Interstellar travel.
Advice for starting a science-driven company: Don’t start too early, don’t spin out too early. Build up as much value as possible before getting diluted and lose some control of your company. At some point, you will make a deal with holders of other IP, so don’t worry too much about IP.
Artificial intelligence foundation for therapeutic science [Nature Chemical Biology, 2022]
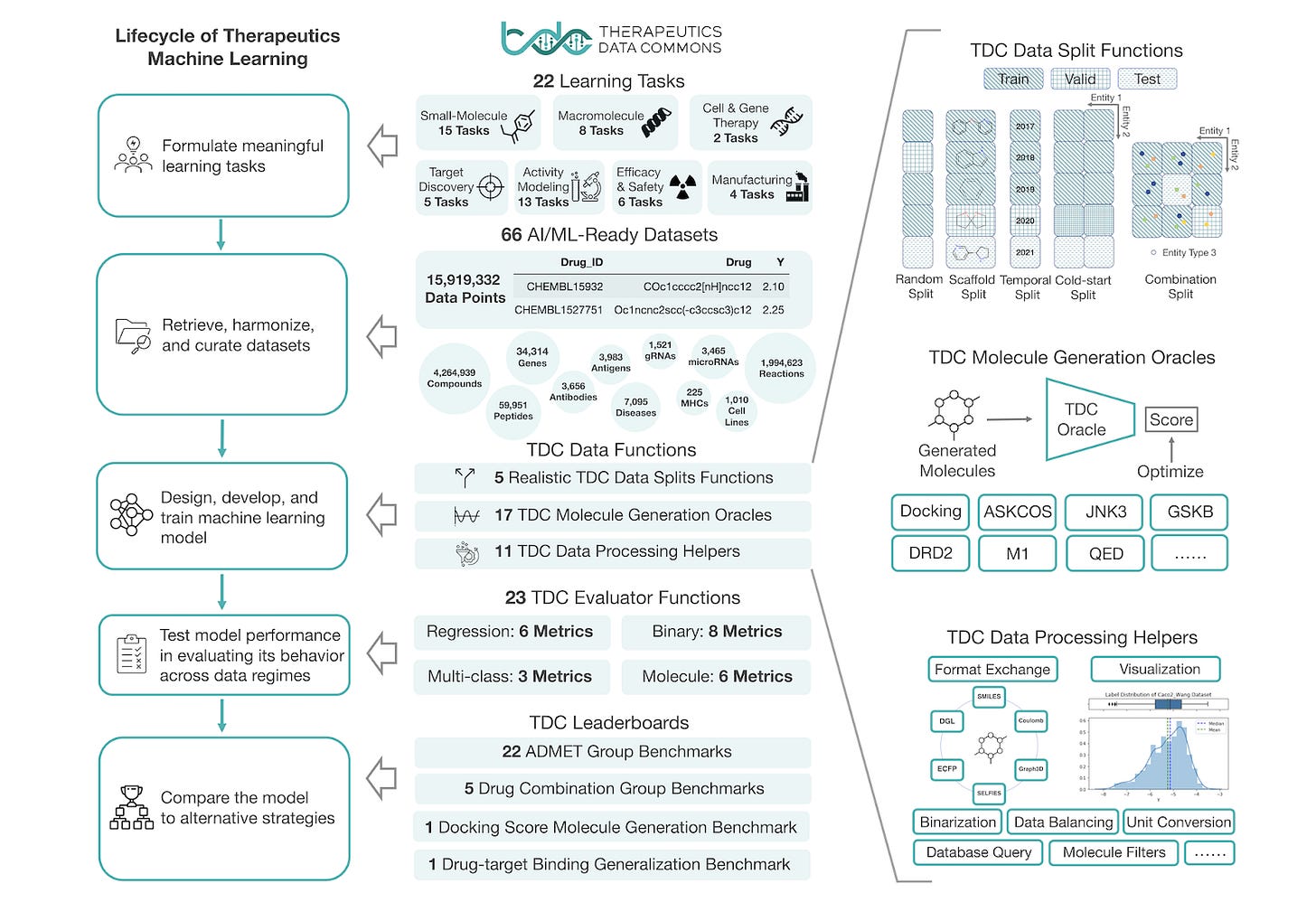
If artificial intelligence and machine learning are going to reach full potential in biotechnology, we need more data. And not just any data. We need more data that is suitable for AI applications and not just the exhaust of what’s happening in the lab. We wanted to elevate the work of Therapeutic Data Commons (aka TDC, thanks Therence!), an open-science machine learning foundation for drug discovery and development.
“At its core, TDC is a collection of AI-solvable tasks, AI-ready datasets and curated benchmarks.” At the time of publication, TDC TDC contains 66 AI-ready datasets that span a total of 15,919,332 data points and are spread across 22 problems in drug discovery including 15 tasks for small molecules such as drug response and synergy prediction, 8 tasks for macromolecules including paratope and epitope prediction, and 2 tasks for cell and gene therapies such as CRISPR repair prediction. In addition to providing benchmarks, method implementations, and implementation tactics for AI in drug discovery, TDC also offers documentation and tutorials.
Why are clinical trials so expensive? Tales from the beast’s belly [MilkyEggs, 2022]
As we’ve previously covered, clinical trial recruitment/logistics are a major obstacle for biotech progress. The author of this piece discusses watching clinical trial management from his/her vantage point as an engineer at a startup running Phase I and Phase II trials. Several issues to clinical trial enrollment and execution are highlighted and almost all point to severe organizational and system-wide inefficiencies. For example, picking a CRO is an unnecessarily lengthy process and involves comparing budgets often based on unsubstantiated estimates. After CROs are picked, focus turns to trial recruitment logistics including patient enrollment, site selection, and data collection. All of which requires specialized pharma knowledge, a wide network, and a lot of back and forth between interested parties (there are no established marketplaces within the clinical trials sphere). Overall, misaligned incentives between companies, clinics, and CROs combined with systemic inefficiencies in decision making lead to unnecessarily inflated trial costs. It will be interesting to see how many of these issues can be alleviated with tech advances vs how many of these issues will require deeper policy and industry-wide change. If you’re looking for an account of one experience with clinical trial organization, this is a fun read.
The 'recalibration phase' of AI in drug discovery
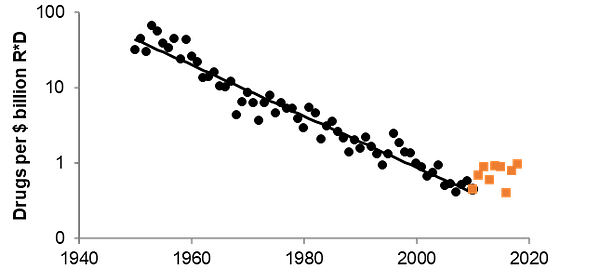
In this piece by Shelby Newsad of Compound VC, we interrogate where the field of techbio is in terms of the traditional “hype cycle.” Grappling with the urgent question of: how do we know if AI/ML approaches are allowing us to traverse novel biochemical space? Are these approaches driving lower costs and greater speed in bringing new therapies to market?
One of our favorite articles featured in this piece by Brender and Cortés-Ciriano showed that despite our advancements in discovery, much of the quality, cost, and speed savings reaped from AI applications in drug discovery and development materials in the clinic during Phase 2. As the article articulates: “...increasing the success of clinical phases decreases the number of expensive clinical trials needed to bring a drug to the market, and this decrease in the number of failures matters more than failing more quickly or more cheaply in terms of cost per successful, approved drug.”
Why is progress in biology so slow? [Samuel G Rodriques, 2022]
When a biotech startup says it’s out to cure a disease, it usually means it wants to find an intervention that will increase lifespan by 3 months. Success usually means an IPO, regardless of whether the drug eventually succeeds or fails
Sam explores what needs to happen to meet the goal of biomedical research: to cure all human disease. We need to solve problems across these three areas:
Speed: it takes 10-20 years to conduct an experiment to solve a human disease and we will be conducting research at snail-pace until we come up with a better ethical and technical framework for faster experimentation. Sam mentions that one of the best uses of machine learning would be to predict drug safety, and subsequently work with regulators to enable drugs that are predicted to be safe with high confidence to be tested directly in humans without pre-clinical trials.
Knowledge: biomedical literature is vast and suffers from unreliability by commission and omission. This means that if you want to know if an experiment really works or what its limitations are, you have to try it out for yourself. Until we have full lab automation, drawing conclusions from existing literature will be unreliable.
Talent: there is no high-status institution like DeepMind or OpenAI for biology. This means that the best talent end up as managers (i.e. as professors or starting a company). We need a new institution and career path that can capture the best talent for biologists to keep doing research
Lecanemab and Alzheimer's: More Data [Derek Lowe, Science, 2022]
I think that both of these are true: the numbers are the best that I have yet seen for such an antibody, and at the same time I doubt if the drug will actually make a difference in the treatment of Alzheimer’s.
Derek opines that it is likely that amyloid is not causative of Alzheimer’s and perhaps a product of the disease, in the same way smoke is a component of a house fire, but not the reason the house burns down.
Eisai and Biogen’s lecanemab has shown the best numbers out of a amyloid-targeting drug. The treatment group in the trial has shown a statistically-significant slow down of CDR-SB (Clinical Dementia Rating - Sum of Boxes), the endpoint and measure used to assess cognition, versus the placebo group. Despite these results, there is still a contentious question around the real-world utility of the drug. Madhav Thambisetty, a neurologist at the National Institute on Aging said “From the perspective of a physician caring for Alzheimer’s patients, the difference between lecanemab and placebo is well below what is considered to be a clinically meaningful treatment effect”. There are also questions around the safety of the drug, with 6.9% patient discontinuation in the treatment group (vs 2.9% of the placebo group).
Ultimately, if lecanemab is so effective at clearing amyloid, yet not great at slowing down the ‘real-world’ progression of the disease, perhaps it is the underlying target that it is not worth much. Given the precedent with aducanumab, a controversial approval by the FDA, it is likely that the agency will have to approve lecanemab as long as it shows sufficient safety.
Academic papers
Illuminating protein space with a programmable generative model [Ingraham et al., Generate Biomedicines preprint, 2022]
Another big week for generative AI for protein engineering. Generate Biomedicines, a Flagship company, published a preprint detailing a generative model called Chroma. The system is built on diffusion models (which we covered in last week’s edition) and graph neural networks. Chroma combines 3 capabilities in one model for the first time:
Generation of full multi-protein complexes
An algorithm that scales sub-quadratically (rather than quadratically) with system size
Programmability via conditional sampling of the model under user specified constraints
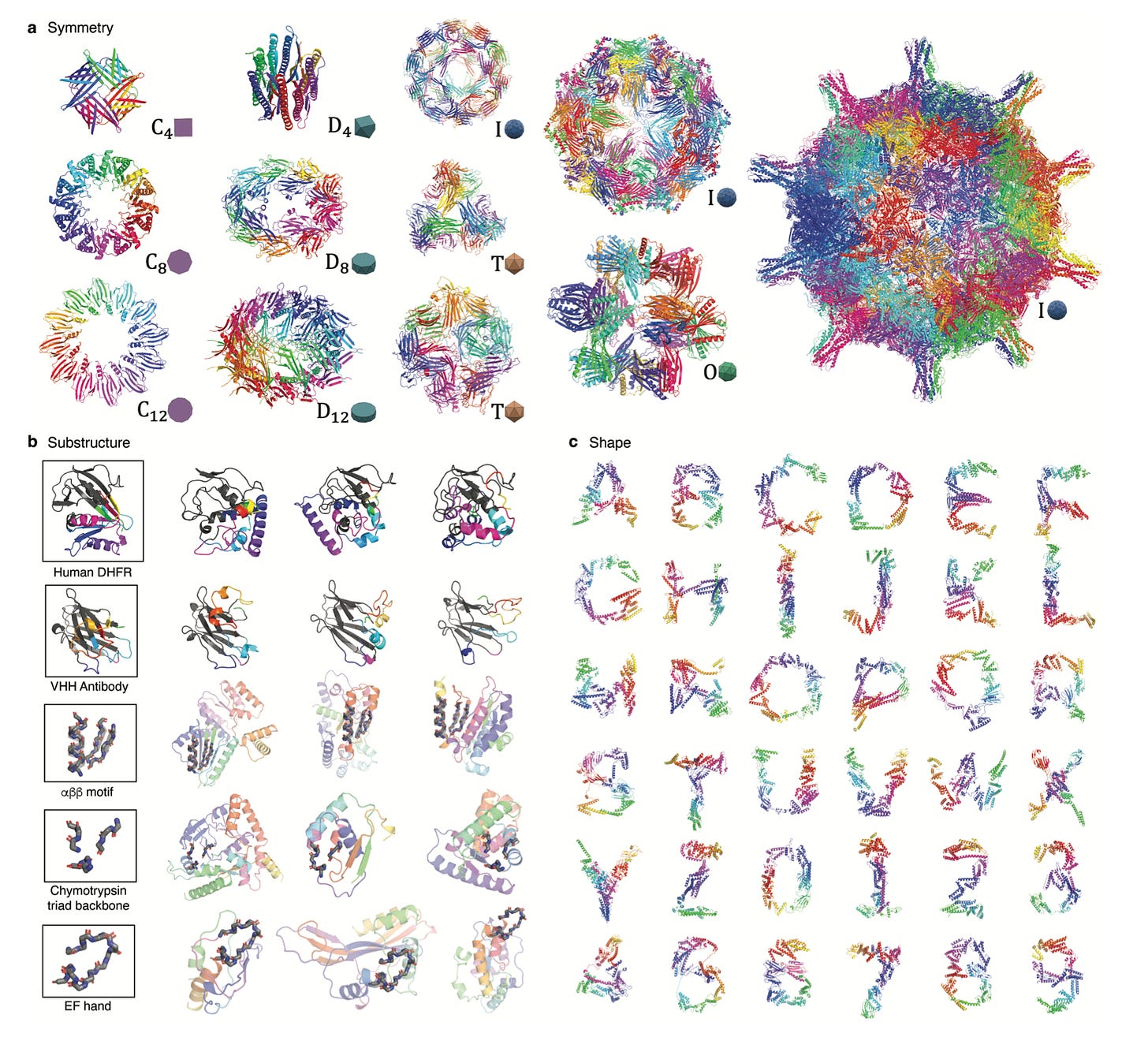
One of the unique features of diffusion models is that for any property X that the algorithm is trained to predict from noisy examples, the model can be repurposed to generate de novo proteins with property X. Leveraging this capability, the model can produce diverse, novel protein structures that are programmable, meaning properties such as symmetry, shape, and protein class can be pre-specified (see figure below for examples).
Why it matters: Computational complexity of previous generative algorithms for protein structure have scaled quadratically in the number of residues, limiting the scalability of the approach for large proteins. Chroma, however, leverages a novel neural network architecture that scales sub-quadratically (interestingly, by taking inspiration from an algorithm for solving the N-body problem in celestial mechanics). Impressively, Chroma can generate extremely large proteins (>3000 residues) on a basic GPU such as an NVIDIA V100 in just a few minutes. There are a couple notable shortcomings. While the authors did some in silico validation of the designability/synthesizability of generated proteins, they did not perform any wet lab validation. The approach does require a fair bit of expert supervision as well; conditional models require tuning and oversight by experienced protein designers for troubleshooting. Regardless, this paper represents a tour de force for de novo protein design.
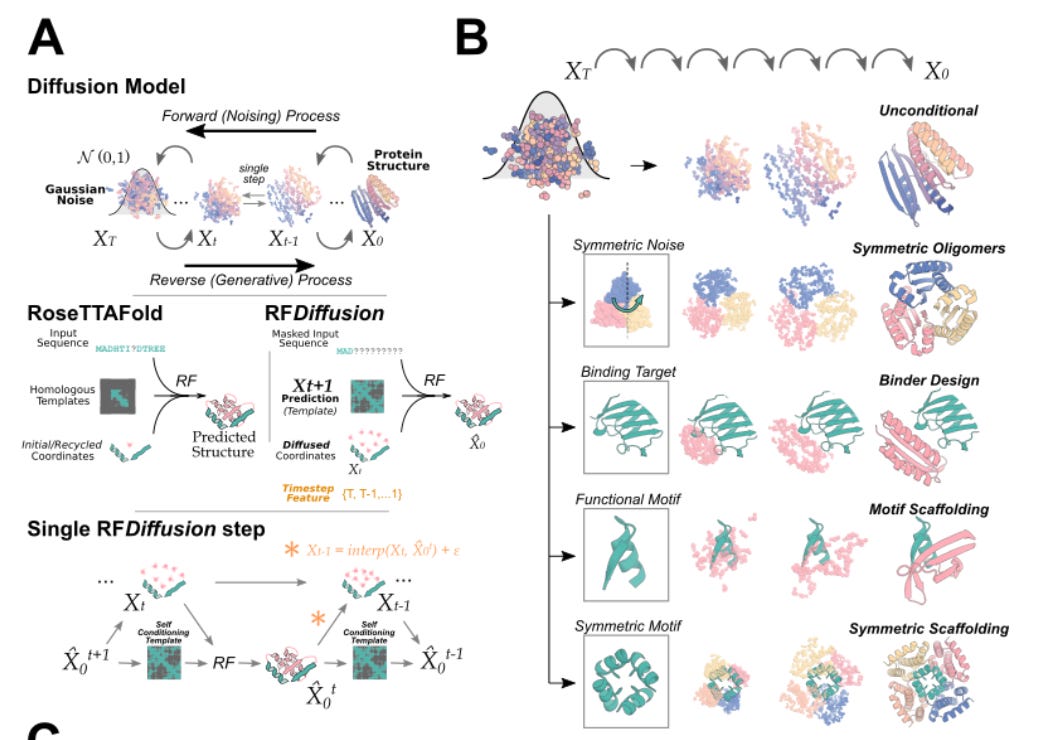
A diffusion model for protein design baker lab [Baker Lab]
A talented team at the Baker Lab has authored a new way of designing proteins by combining structure prediction networks and generative diffusion models. We’ve talked about our excitement for diffusion models in the past, which at the highest level, is a machine learning method that is adeptly skilled at adding and removing noise from data assets to generate new, plausible content. DALL-E should ring a bell as a diffusion model that can generate new images. The team at the Baker Lab created a new method called RF Diffusion that can generate novel proteins that bind to molecular targets and be configured to produce symmetric or asymmetric oligomers, building upon existing protein structure prediction networks like RoseTTAFold.
Why it matters: In a brilliantly combinatorial approach, RF Diffusion leverages RoseTTAFOLD as the denoising network in a generative diffusion model, dramatically increasing the protein structure space explored by deterministic approaches. Importantly, “RFdiffusion readily generates complex protein structures with little overall structural similarity to any known protein structures, indicating considerable generalization beyond the PDB training set.” In building upon foundation models such as existing protein structure prediction methods, this work underscores the immense opportunity in combinatorial computational approaches to improve performance and address determinism in existing methods.Check out the pre-pre-print here, soon to be published on BioRxiv. Similar work was published by Generate Biomedicines described above.
Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases [Yarnall et al., Nature Biotechnology, 2022]
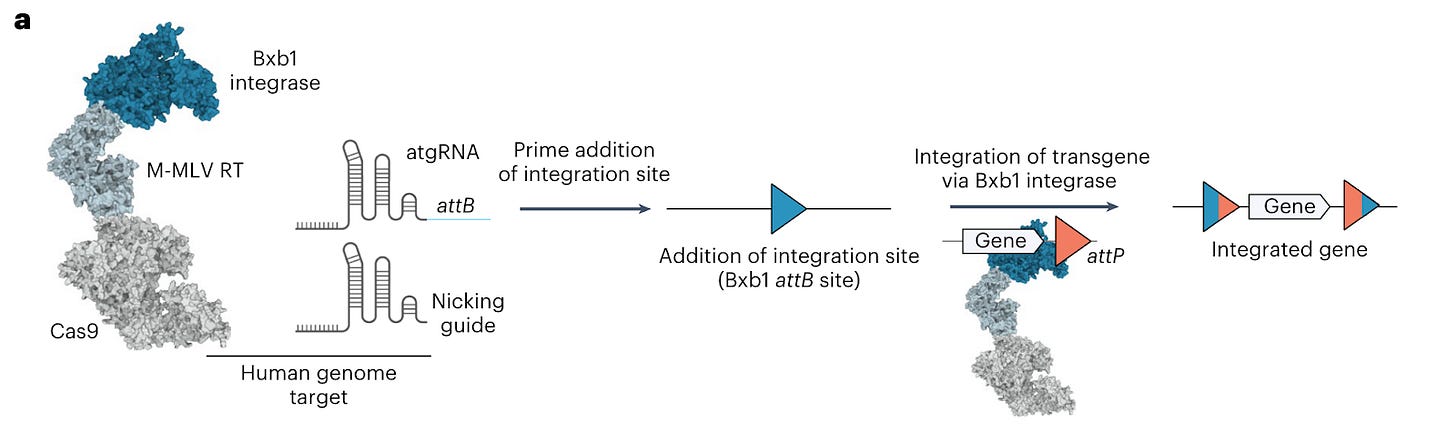
We’re probably too excited about this paper. Omar Abudayyeh, Jonathan Gootenberg and the rest of the team present programmable addition via site-specific targeting elements (PASTE), which uses CRISPR-Cas9 nickase fused to both a reverse transcriptase (as with prime editing) and a serine integrase for payload integration.
Why it matters: PASTE allows large, multiplexed gene insertion without reliance on DNA repair pathways. Common gene editing methods such as CRISPR-Cas9 rely on double-strand breaks (DSBs) of DNA and endogenous DNA repair mechanisms to facilitate the integration of new genes. However, DSB-based approaches have limitations as it can cause unintended insertions/deletions, translocations and p53 activation which can cause damage. Other approaches such as base editing and prime editing can alleviate issues caused by DSBs, but they are limited to nucleotide edits or small insertions. This can enable the therapeutic correction of genetic disease for recessive loss of function mutations and overcoming dominant-negative mutations. Current approaches for such diseases are mutation-sensitive, whereas PASTE could be mutation-agnostic and thus targeting a wider patient population per therapy.The authors demonstrated that PASTE can insert cargos of up to 36 kB in a single delivery with efficiencies of up to ~50-60% in cell lines and ~4-5% in primary human hepatocytes and T-cells. The PASTE system (see Figure A) involves insertion of landing sites via Cas-9 directed reverse transcriptases (pegRNAs containing attB sequences, named atgRNA), followed by landing site recognition and integration of cargo via Cas9-directed integrases (containing the attP attachment site).
According to Endpoints, the technology has been licensed to Tome Biosciences — a biotech co-founded by Abudayyeh and Gootenbergin in 2021 and backed by ARCH, GV, a16z, Longwood Fund, Polaris Partners and Alexandria Venture.
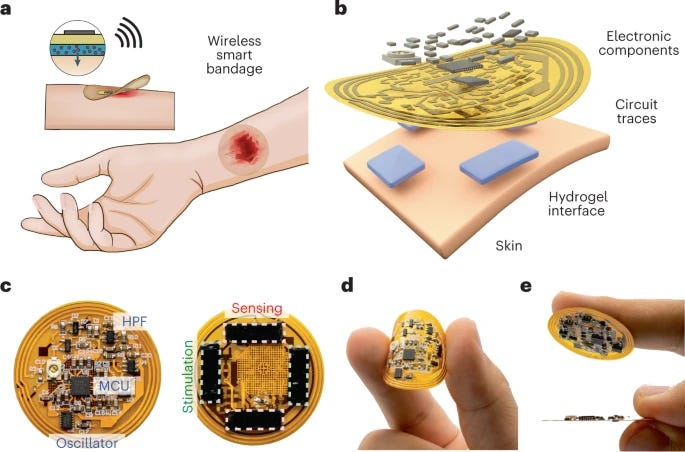
Wireless, closed-loop, smart bandage with integrated sensors and stimulators for advanced wound care and accelerated healing [Jiang et al., Nature Biotechnology, 2022]
Zhenan Bao and team strikes again – pushing the envelope forward in the realm of flexible electronics and their application to medicine.
Why it matters: Chronic non-healing wounds are a tremendous medical burden, affecting over 6M+ people in the U.S. alone, with costs exceeding $25B annually. Such wounds cascade into a series of psychological and physical complications – such as disability, depression, and prolonged hospitalization – with overall increased morbidity and mortality. Current wound dressings are simple (i.e. cover a la bandage) and do not respond to the healing wound environment, which is dynamic and complex. In this paper, the authors develop a wireless smart bandage that integrates sensory and treatment modalities for real-time monitoring and active wound care with minimal clinician engagement. The authors developed a miniature flexible PCB that was capable of sensing 1) wound impedance (skin to electric current - a marker of how healed the wound is) and 2) temperature ( ↑ temp = ↑ likelihood of infection) while 3) delivering programmed electrical stimulation. In preclinical mouse models, this smart band aid could successfully monitor wounds and facilitate increased neovascularization and enhanced dermal recovery. Their data suggested that electrical stimulation may drive macrophages towards a pro-regenerative phenotype, which may be responsive to local microenvironmental stimuli.
What we listened to
Machine Learning in Structural Biology Workshop [NeurIPS]
Watch here.
The Antibody Engine [Business of Biotech]
Model Term Sheets, Pragmatic Trials, & Gene Therapy Prices [BioCentury This Week]
Notable Deals
Entact Bio makes its Series A debut with $81M to launch protein enhancement platform.
Entact is developing the Encompass deubiquitinase platform for stabilizing proteins. These enhancement-targeting chimeric (ENTAC) small molecules will be used in diseases where patients have too little of a protein in cells, with the goal of enhancing or stabilizing them. The round was led by Qiming Venture Partners USA and venBio Partners, with other investors including Abingworth, Brandon Capital, Janus Henderson Investors, Logos Capital, Surveyor Capital and WEHI. Founding investors were 4BIO Capital and Arkin Bio Ventures.
Intellia and Iveric sell stocks to raise money, each netting $300M.
Intellia’s raise is just a few weeks after positive data from their hereditary angioedema program. The $300M will be used to advance Intellia’s pre-clinical and clinical CRISPR-based therapeutics. Iveric is currently working towards an FDA submission for its drug targeted geographic atrophy (AMD); this money would be used to push forward this drug, avacincaptad pegol, a complement C5 protein inhibitor.
Bionaut Labs gets $43.2M for its tiny drug delivery robots
Delivery, delivery delivery. Bionaut is developing magnetically-driven drug delivery robots that are designed to deliver treatments to the midbrain - going after a number of central nervous system pathologies. This round was led by Khosla Ventures and featured new investors Deep Insight, OurCrowd, PSPRS, Sixty Degree Capital, Dolby Family Ventures, GISEV Family Ventures, what if ventures, Tintah Grace and Gaingels. The Series A capital came from YFM, Maven, Triple Point Ventures, Northern Powerhouse Investment Fund and Manchester Tech Trust angels.
AI drug discovery startup Biorelate bags £6.5m ahead of US move
Biorelate’s Galactic AI platform uses NLP and ML to collect biomedical data from any source of text for downstream drug-development. Each text article runs through a series of deep learning and natural language processing software services that have been built to understand biomedical research. Millions of articles are routinely auto-curated in just under 6 hours.
Rallybio, AbCellera form new partnership around antibodies for rare disease
Both biotechs will co-develop drugs against up to 5 rare disease targets; this will combine AbCellera’s antibody discovery engine with Rallybio’s expertise in rare disease. This is on the heels of a partnership with Moderna last year, on up to six undisclosed targets for mRNA-encoded therapeutics.
What we liked on Twitter





Field Trip



Congratulations to the entire Lux Capital team on their continued success + additions to the team!
Did we miss anything? Would you like to contribute to Decoding TechBio by writing a guest post? Drop us a note here or chat with us on Twitter: @ameekapadia @pablolubroth @patricksmalone @morgancheatham @ketanyerneni
















These posts are excellent, week in and week out. Love the "Why It Matters" boxes.